Crystal structures of Leptospira interrogans FAD-containing ferredoxin-NADP+ reductase and its complex with NADP+
- PMID: 17958910
- PMCID: PMC2148051
- DOI: 10.1186/1472-6807-7-69
Crystal structures of Leptospira interrogans FAD-containing ferredoxin-NADP+ reductase and its complex with NADP+
Abstract
Background: Ferredoxin-NADP(H) reductases (FNRs) are flavoenzymes that catalyze the electron transfer between NADP(H) and the proteins ferredoxin or flavodoxin. A number of structural features distinguish plant and bacterial FNRs, one of which is the mode of the cofactor FAD binding. Leptospira interrogans is a spirochaete parasitic bacterium capable of infecting humans and mammals in general. Leptospira interrogans FNR (LepFNR) displays low sequence identity with plant (34% with Zea mays) and bacterial (31% with Escherichia coli) FNRs. However, LepFNR contains all consensus sequences that define the plastidic class FNRs.
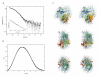
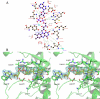
Results: The crystal structures of the FAD-containing LepFNR and the complex of the enzyme with NADP+, were solved and compared to known FNRs. The comparison reveals significant structural similarities of the enzyme with the plastidic type FNRs and differences with the bacterial enzymes. Our small angle X-ray scattering experiments show that LepFNR is a monomeric enzyme. Moreover, our biochemical data demonstrate that the LepFNR has an enzymatic activity similar to those reported for the plastidic enzymes and that is significantly different from bacterial flavoenzymes, which display lower turnover rates.
Conclusion: LepFNR is the first plastidic type FNR found in bacteria and, despite of its low sequence similarity with plastidic FNRs still displays high catalytic turnover rates. The typical structural and biochemical characteristics of plant FNRs unveiled for LepFNR support a notion of a putative lateral gene transfer which presumably offers Leptospira interrogans evolutionary advantages. The wealth of structural information about LepFNR provides a molecular basis for advanced drugs developments against leptospirosis.
Figures


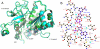
Similar articles
-
A highly stable plastidic-type ferredoxin-NADP(H) reductase in the pathogenic bacterium Leptospira interrogans.PLoS One. 2011;6(10):e26736. doi: 10.1371/journal.pone.0026736. Epub 2011 Oct 24. PLoS One. 2011. PMID: 22039544 Free PMC article.
-
A bacterial 2[4Fe4S] ferredoxin as redox partner of the plastidic-type ferredoxin-NADP+ reductase from Leptospira interrogans.Biochim Biophys Acta Gen Subj. 2019 Apr;1863(4):651-660. doi: 10.1016/j.bbagen.2019.01.004. Epub 2019 Jan 11. Biochim Biophys Acta Gen Subj. 2019. PMID: 30639162
-
Swapping FAD binding motifs between plastidic and bacterial ferredoxin-NADP(H) reductases.Biochemistry. 2011 Mar 29;50(12):2111-22. doi: 10.1021/bi101772a. Epub 2011 Mar 2. Biochemistry. 2011. PMID: 21306142
-
Functional plasticity and catalytic efficiency in plant and bacterial ferredoxin-NADP(H) reductases.Biochim Biophys Acta. 2004 May 6;1698(2):155-65. doi: 10.1016/j.bbapap.2003.12.005. Biochim Biophys Acta. 2004. PMID: 15134648 Review.
-
Structural and functional diversity of ferredoxin-NADP(+) reductases.Arch Biochem Biophys. 2008 Jun 15;474(2):283-91. doi: 10.1016/j.abb.2008.02.014. Epub 2008 Feb 16. Arch Biochem Biophys. 2008. PMID: 18307973 Review.
Cited by
-
A highly stable plastidic-type ferredoxin-NADP(H) reductase in the pathogenic bacterium Leptospira interrogans.PLoS One. 2011;6(10):e26736. doi: 10.1371/journal.pone.0026736. Epub 2011 Oct 24. PLoS One. 2011. PMID: 22039544 Free PMC article.
-
Structural-functional characterization and physiological significance of ferredoxin-NADP reductase from Xanthomonas axonopodis pv. citri.PLoS One. 2011;6(11):e27124. doi: 10.1371/journal.pone.0027124. Epub 2011 Nov 9. PLoS One. 2011. PMID: 22096528 Free PMC article.
-
Ferredoxin-NADP+ reductase from Pseudomonas putida functions as a ferric reductase.J Bacteriol. 2009 Mar;191(5):1472-9. doi: 10.1128/JB.01473-08. Epub 2008 Dec 29. J Bacteriol. 2009. PMID: 19114475 Free PMC article.
-
Structural and mutational analyses of the Leptospira interrogans virulence-related heme oxygenase provide insights into its catalytic mechanism.PLoS One. 2017 Aug 3;12(8):e0182535. doi: 10.1371/journal.pone.0182535. eCollection 2017. PLoS One. 2017. PMID: 28771589 Free PMC article.
References
-
- Ceccarelli EA, Arakaki AK, Cortez N, Carrillo N. Functional plasticity and catalytic efficiency in plant and bacterial ferredoxin-NADP(H) reductases. Biochim Biophys Acta. 2004;1968:155–165. - PubMed
-
- Deng Z, Aliverti A, Zanetti G, Araraki AK, Ottado J, Orellano EG, Calcaterra NB, Ceccarelli EA, Carrillo N, Karplus PA. A productive NADP+ binding mode of ferredoxin-NADP+ reductase revealed by protein engineering and crystallographic studies. Nat Struct Biol. 1999;6:847–853. doi: 10.1038/12307. - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

