EuCAP, a Eukaryotic Community Annotation Package, and its application to the rice genome
- PMID: 17961238
- PMCID: PMC2151081
- DOI: 10.1186/1471-2164-8-388
EuCAP, a Eukaryotic Community Annotation Package, and its application to the rice genome
Abstract
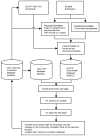
Background: Despite the improvements of tools for automated annotation of genome sequences, manual curation at the structural and functional level can provide an increased level of refinement to genome annotation. The Institute for Genomic Research Rice Genome Annotation (hereafter named the Osa1 Genome Annotation) is the product of an automated pipeline and, for this reason, will benefit from the input of biologists with expertise in rice and/or particular gene families. Leveraging knowledge from a dispersed community of scientists is a demonstrated way of improving a genome annotation. This requires tools that facilitate 1) the submission of gene annotation to an annotation project, 2) the review of the submitted models by project annotators, and 3) the incorporation of the submitted models in the ongoing annotation effort.
Results: We have developed the Eukaryotic Community Annotation Package (EuCAP), an annotation tool, and have applied it to the rice genome. The primary level of curation by community annotators (CA) has been the annotation of gene families. Annotation can be submitted by email or through the EuCAP Web Tool. The CA models are aligned to the rice pseudomolecules and the coordinates of these alignments, along with functional annotation, are stored in the MySQL EuCAP Gene Model database. Web pages displaying the alignments of the CA models to the Osa1 Genome models are automatically generated from the EuCAP Gene Model database. The alignments are reviewed by the project annotators (PAs) in the context of experimental evidence. Upon approval by the PAs, the CA models, along with the corresponding functional annotations, are integrated into the Osa1 Genome Annotation. The CA annotations, grouped by family, are displayed on the Community Annotation pages of the project website http://rice.tigr.org, as well as in the Community Annotation track of the Genome Browser.
Conclusion: We have applied EuCAP to rice. As of July 2007, the structural and/or functional annotation of 1,094 genes representing 57 families have been deposited and integrated into the current gene set. All of the EuCAP components are open-source, thereby allowing the implementation of EuCAP for the annotation of other genomes. EuCAP is available at http://sourceforge.net/projects/eucap/.
Figures


Similar articles
-
The institute for genomic research Osa1 rice genome annotation database.Plant Physiol. 2005 May;138(1):18-26. doi: 10.1104/pp.104.059063. Plant Physiol. 2005. PMID: 15888674 Free PMC article.
-
The TIGR Rice Genome Annotation Resource: improvements and new features.Nucleic Acids Res. 2007 Jan;35(Database issue):D883-7. doi: 10.1093/nar/gkl976. Epub 2006 Dec 1. Nucleic Acids Res. 2007. PMID: 17145706 Free PMC article.
-
The TIGR rice genome annotation resource: annotating the rice genome and creating resources for plant biologists.Nucleic Acids Res. 2003 Jan 1;31(1):229-33. doi: 10.1093/nar/gkg059. Nucleic Acids Res. 2003. PMID: 12519988 Free PMC article.
-
Annotating the genome of Medicago truncatula.Curr Opin Plant Biol. 2006 Apr;9(2):122-7. doi: 10.1016/j.pbi.2006.01.004. Epub 2006 Feb 2. Curr Opin Plant Biol. 2006. PMID: 16458040 Review.
-
From plant genomes to protein families: computational tools.Comput Struct Biotechnol J. 2013 Aug 14;8:e201307001. doi: 10.5936/csbj.201307001. eCollection 2013. Comput Struct Biotechnol J. 2013. PMID: 24688740 Free PMC article. Review.
Cited by
-
Genomic and genetic database resources for the grasses.Plant Physiol. 2009 Jan;149(1):132-6. doi: 10.1104/pp.108.129593. Plant Physiol. 2009. PMID: 19126704 Free PMC article. Review. No abstract available.
-
Identification and characterization of pseudogenes in the rice gene complement.BMC Genomics. 2009 Jul 16;10:317. doi: 10.1186/1471-2164-10-317. BMC Genomics. 2009. PMID: 19607679 Free PMC article.
-
An improved genome release (version Mt4.0) for the model legume Medicago truncatula.BMC Genomics. 2014 Apr 27;15:312. doi: 10.1186/1471-2164-15-312. BMC Genomics. 2014. PMID: 24767513 Free PMC article.
References
-
- Haas BJ, Delcher AL, Mount SM, Wortman JR, Smith RK, Jr., Hannick LI, Maiti R, Ronning CM, Rusch DB, Town CD, Salzberg SL, White O. Improving the Arabidopsis genome annotation using maximal transcript alignment assemblies. Nucleic Acids Res. 2003;31:5654–5666. doi: 10.1093/nar/gkg770. - DOI - PMC - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Research Materials

