Large-scale analysis by SAGE reveals new mechanisms of v-erbA oncogene action
- PMID: 17961265
- PMCID: PMC2194726
- DOI: 10.1186/1471-2164-8-390
Large-scale analysis by SAGE reveals new mechanisms of v-erbA oncogene action
Abstract
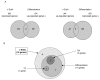
Background: The v-erbA oncogene, carried by the Avian Erythroblastosis Virus, derives from the c-erbAalpha proto-oncogene that encodes the nuclear receptor for triiodothyronine (T3R). v-ErbA transforms erythroid progenitors in vitro by blocking their differentiation, supposedly by interference with T3R and RAR (Retinoic Acid Receptor). However, v-ErbA target genes involved in its transforming activity still remain to be identified.
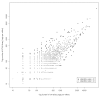
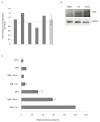
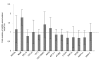
Results: By using Serial Analysis of Gene Expression (SAGE), we identified 110 genes deregulated by v-ErbA and potentially implicated in the transformation process. Bioinformatic analysis of promoter sequence and transcriptional assays point out a potential role of c-Myb in the v-ErbA effect. Furthermore, grouping of newly identified target genes by function revealed both expected (chromatin/transcription) and unexpected (protein metabolism) functions potentially deregulated by v-ErbA. We then focused our study on 15 of the new v-ErbA target genes and demonstrated by real time PCR that in majority their expression was activated neither by T3, nor RA, nor during differentiation. This was unexpected based upon the previously known role of v-ErbA.
Conclusion: This paper suggests the involvement of a wealth of new unanticipated mechanisms of v-ErbA action.
Figures



References
-
- Graf T, Beug H. Avian leukemia viruses: interaction with their target cells in vivo and in vitro. Biochim Biophys Acta. 1978;516:269–299. - PubMed
-
- Casini T, Graf T. Bicistronic retroviral vector reveals capacity of v-erbA to induce erythroleukemia and to co-operate with v-myb. Oncogene. 1995;11:1019–1026. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

