Expression of a novel non-coding mitochondrial RNA in human proliferating cells
- PMID: 17962305
- PMCID: PMC2175360
- DOI: 10.1093/nar/gkm863
Expression of a novel non-coding mitochondrial RNA in human proliferating cells
Abstract
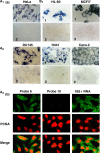
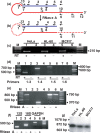
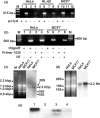
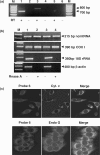
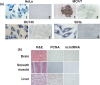
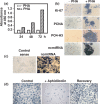
Previously, we reported the presence in mouse cells of a mitochondrial RNA which contains an inverted repeat (IR) of 121 nucleotides (nt) covalently linked to the 5' end of the mitochondrial 16S RNA (16S mtrRNA). Here, we report the structure of an equivalent transcript of 2374 nt which is over-expressed in human proliferating cells but not in resting cells. The transcript contains a hairpin structure comprising an IR of 815 nt linked to the 5' end of the 16S mtrRNA and forming a long double-stranded structure or stem and a loop of 40 nt. The stem is resistant to RNase A and can be detected and isolated after digestion with the enzyme. This novel transcript is a non-coding RNA (ncRNA) and several evidences suggest that the transcript is synthesized in mitochondria. The expression of this transcript can be induced in resting lymphocytes stimulated with phytohaemagglutinin (PHA). Moreover, aphidicolin treatment of DU145 cells reversibly blocks proliferation and expression of the transcript. If the drug is removed, the cells re-assume proliferation and over-express the ncmtRNA. These results suggest that the expression of the ncmtRNA correlates with the replicative state of the cell and it may play a role in cell proliferation.
Figures

Similar articles
-
Expression of a family of noncoding mitochondrial RNAs distinguishes normal from cancer cells.Proc Natl Acad Sci U S A. 2009 Jun 9;106(23):9430-4. doi: 10.1073/pnas.0903086106. Epub 2009 May 26. Proc Natl Acad Sci U S A. 2009. PMID: 19470459 Free PMC article.
-
Coding RNAs with a non-coding function: maintenance of open chromatin structure.Nucleus. 2011 Sep-Oct;2(5):410-24. doi: 10.4161/nucl.2.5.17736. Epub 2011 Sep 1. Nucleus. 2011. PMID: 21983088
-
A putative RNA editing from U to C in a mouse mitochondrial transcript.Nucleic Acids Res. 2002 May 1;30(9):1895-901. doi: 10.1093/nar/30.9.1895. Nucleic Acids Res. 2002. PMID: 11972325 Free PMC article.
-
Non-coding-RNA regulators of RNA polymerase II transcription.Nat Rev Mol Cell Biol. 2006 Aug;7(8):612-6. doi: 10.1038/nrm1946. Epub 2006 May 24. Nat Rev Mol Cell Biol. 2006. PMID: 16723972 Review.
-
Long non-coding RNAs: insights into functions.Nat Rev Genet. 2009 Mar;10(3):155-9. doi: 10.1038/nrg2521. Nat Rev Genet. 2009. PMID: 19188922 Review.
Cited by
-
Nuclear localization of the mitochondrial ncRNAs in normal and cancer cells.Cell Oncol (Dordr). 2011 Aug;34(4):297-305. doi: 10.1007/s13402-011-0018-8. Epub 2011 Feb 24. Cell Oncol (Dordr). 2011. PMID: 21347712
-
Role of lncRNAs in Cellular Aging.Front Endocrinol (Lausanne). 2016 Dec 6;7:151. doi: 10.3389/fendo.2016.00151. eCollection 2016. Front Endocrinol (Lausanne). 2016. PMID: 27999563 Free PMC article. Review.
-
Long noncoding RNAs coordinate functions between mitochondria and the nucleus.Epigenetics Chromatin. 2017 Aug 23;10(1):41. doi: 10.1186/s13072-017-0149-x. Epigenetics Chromatin. 2017. PMID: 28835257 Free PMC article. Review.
-
Mitochondrial Gene Expression and Beyond-Novel Aspects of Cellular Physiology.Cells. 2019 Dec 19;9(1):17. doi: 10.3390/cells9010017. Cells. 2019. PMID: 31861673 Free PMC article. Review.
-
Mitochondrial Non-Coding RNAs Are Potential Mediators of Mitochondrial Homeostasis.Biomolecules. 2022 Dec 13;12(12):1863. doi: 10.3390/biom12121863. Biomolecules. 2022. PMID: 36551291 Free PMC article. Review.
References
-
- Nass MM, Nass S, Afzelius BA. The general occurrence of mitochondrial DNA. Exp. Cell Res. 1965;37:516–539. - PubMed
-
- Anderson S, Bankier AT, Barrell BG, de Brujin MH, Coulson AR, Drouin J, Eperon IC, Nierlich DP, Roe BA, et al. Sequence and organization of the human mitochondrial genome. Nature. 1981;290:457–465. - PubMed
-
- Ojala D, Montoya J, Attardi G. tRNA punctuation model of RNA processing in human mitochondria. Nature. 1981;290:470–474. - PubMed
-
- Chomyn A, Attardi G. Recent advances on mitochondrial biogenesis. In: Ernster L, editor. Molecular Mechanisms in Biogenesis. Amsterdam: Elsevier Science Publishers B. V.; 1992. pp. 483–509.
-
- Shadel GS, Clayton DA. Mitochondrial DNA maintenance in vertebrates. Ann. Rev. Biochem. 1997;66:409–435. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

