ProCKSI: a decision support system for Protein (structure) Comparison, Knowledge, Similarity and Information
- PMID: 17963510
- PMCID: PMC2222653
- DOI: 10.1186/1471-2105-8-416
ProCKSI: a decision support system for Protein (structure) Comparison, Knowledge, Similarity and Information
Abstract
Background: We introduce the decision support system for Protein (Structure) Comparison, Knowledge, Similarity and Information (ProCKSI). ProCKSI integrates various protein similarity measures through an easy to use interface that allows the comparison of multiple proteins simultaneously. It employs the Universal Similarity Metric (USM), the Maximum Contact Map Overlap (MaxCMO) of protein structures and other external methods such as the DaliLite and the TM-align methods, the Combinatorial Extension (CE) of the optimal path, and the FAST Align and Search Tool (FAST). Additionally, ProCKSI allows the user to upload a user-defined similarity matrix supplementing the methods mentioned, and computes a similarity consensus in order to provide a rich, integrated, multicriteria view of large datasets of protein structures.
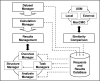
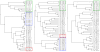
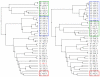
Results: We present ProCKSI's architecture and workflow describing its intuitive user interface, and show its potential on three distinct test-cases. In the first case, ProCKSI is used to evaluate the results of a previous CASP competition, assessing the similarity of proposed models for given targets where the structures could have a large deviation from one another. To perform this type of comparison reliably, we introduce a new consensus method. The second study deals with the verification of a classification scheme for protein kinases, originally derived by sequence comparison by Hanks and Hunter, but here we use a consensus similarity measure based on structures. In the third experiment using the Rost and Sander dataset (RS126), we investigate how a combination of different sets of similarity measures influences the quality and performance of ProCKSI's new consensus measure. ProCKSI performs well with all three datasets, showing its potential for complex, simultaneous multi-method assessment of structural similarity in large protein datasets. Furthermore, combining different similarity measures is usually more robust than relying on one single, unique measure.
Conclusion: Based on a diverse set of similarity measures, ProCKSI computes a consensus similarity profile for the entire protein set. All results can be clustered, visualised, analysed and easily compared with each other through a simple and intuitive interface.ProCKSI is publicly available at http://www.procksi.net for academic and non-commercial use.
Figures

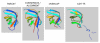

Similar articles
-
Toward high-throughput, multicriteria protein-structure comparison and analysis.IEEE Trans Nanobioscience. 2010 Jun;9(2):144-55. doi: 10.1109/TNB.2010.2043851. IEEE Trans Nanobioscience. 2010. PMID: 20650704
-
Compression-based classification of biological sequences and structures via the Universal Similarity Metric: experimental assessment.BMC Bioinformatics. 2007 Jul 13;8:252. doi: 10.1186/1471-2105-8-252. BMC Bioinformatics. 2007. PMID: 17629909 Free PMC article.
-
Classification of rhodopsin structures by modern methods of structural bioinformatics.Biochemistry (Mosc). 2012 May;77(5):435-43. doi: 10.1134/S0006297912050033. Biochemistry (Mosc). 2012. PMID: 22813584
-
Measuring the similarity of protein structures by means of the universal similarity metric.Bioinformatics. 2004 May 1;20(7):1015-21. doi: 10.1093/bioinformatics/bth031. Epub 2004 Jan 29. Bioinformatics. 2004. PMID: 14751983
-
Folic acid supplementation and malaria susceptibility and severity among people taking antifolate antimalarial drugs in endemic areas.Cochrane Database Syst Rev. 2022 Feb 1;2(2022):CD014217. doi: 10.1002/14651858.CD014217. Cochrane Database Syst Rev. 2022. PMID: 36321557 Free PMC article.
Cited by
-
A simple and fast heuristic for protein structure comparison.BMC Bioinformatics. 2008 Mar 25;9:161. doi: 10.1186/1471-2105-9-161. BMC Bioinformatics. 2008. PMID: 18366735 Free PMC article.
-
Efficient Multicriteria Protein Structure Comparison on Modern Processor Architectures.Biomed Res Int. 2015;2015:563674. doi: 10.1155/2015/563674. Epub 2015 Oct 28. Biomed Res Int. 2015. PMID: 26605332 Free PMC article.
-
Functional and structural characterization of a thermostable acetyl esterase from Thermotoga maritima.Proteins. 2012 Jun;80(6):1545-59. doi: 10.1002/prot.24041. Epub 2012 Mar 13. Proteins. 2012. PMID: 22411095 Free PMC article.
-
CSA: comprehensive comparison of pairwise protein structure alignments.Nucleic Acids Res. 2012 Jul;40(Web Server issue):W303-9. doi: 10.1093/nar/gks362. Epub 2012 May 2. Nucleic Acids Res. 2012. PMID: 22553365 Free PMC article.
-
Fast Phylogeny of SARS-CoV-2 by Compression.Entropy (Basel). 2022 Mar 22;24(4):439. doi: 10.3390/e24040439. Entropy (Basel). 2022. PMID: 35455102 Free PMC article.
References
-
- Ferro D, Hermans J. A Different Best Rigid-body Molecular Fit Routine. Acta Crystallogr. 1977;A33:345–347.
-
- Kabsch W. A Discussion of the Solution for the Best Rotation to Relate Two Sets of Vectors. Acta Crystallogr. 1978;A34:827–828.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

