HMGB binding to DNA: single and double box motifs
- PMID: 17964600
- PMCID: PMC2117627
- DOI: 10.1016/j.jmb.2007.09.073
HMGB binding to DNA: single and double box motifs
Erratum in
- J Mol Biol. 2009 Mar 27;387(2):520
Abstract
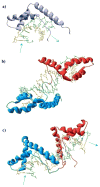
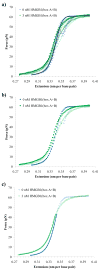
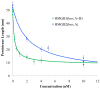
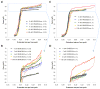
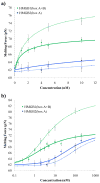
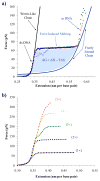
High mobility group (HMG) proteins are nuclear proteins believed to significantly affect DNA interactions by altering nucleic acid flexibility. Group B (HMGB) proteins contain HMG box domains known to bind to the DNA minor groove without sequence specificity, slightly intercalating base pairs and inducing a strong bend in the DNA helical axis. A dual-beam optical tweezers system is used to extend double-stranded DNA (dsDNA) in the absence as well as presence of a single box derivative of human HMGB2 [HMGB2(box A)] and a double box derivative of rat HMGB1 [HMGB1(box A+box B)]. The single box domain is observed to reduce the persistence length of the double helix, generating sharp DNA bends with an average bending angle of 99+/-9 degrees and, at very high concentrations, stabilizing dsDNA against denaturation. The double box protein contains two consecutive HMG box domains joined by a flexible tether. This protein also reduces the DNA persistence length, induces an average bending angle of 77+/-7 degrees , and stabilizes dsDNA at significantly lower concentrations. These results suggest that single and double box proteins increase DNA flexibility and stability, albeit both effects are achieved at much lower protein concentrations for the double box. In addition, at low concentrations, the single box protein can alter DNA flexibility without stabilizing dsDNA, whereas stabilization at higher concentrations is likely achieved through a cooperative binding mode.
Figures
Similar articles
-
Dual binding modes for an HMG domain from human HMGB2 on DNA.Biophys J. 2005 Jul;89(1):353-64. doi: 10.1529/biophysj.104.052068. Epub 2005 Apr 15. Biophys J. 2005. PMID: 15833996 Free PMC article.
-
Transient HMGB protein interactions with B-DNA duplexes and complexes.Biochem Biophys Res Commun. 2008 Jun 20;371(1):79-84. doi: 10.1016/j.bbrc.2008.04.024. Epub 2008 Apr 14. Biochem Biophys Res Commun. 2008. PMID: 18413230 Free PMC article.
-
Enhancement of DNA flexibility in vitro and in vivo by HMGB box A proteins carrying box B residues.Biochemistry. 2009 Mar 17;48(10):2125-34. doi: 10.1021/bi802269f. Biochemistry. 2009. PMID: 19236006 Free PMC article.
-
HMGB proteins and arthritis.Hum Cell. 2018 Jan;31(1):1-9. doi: 10.1007/s13577-017-0182-x. Epub 2017 Sep 15. Hum Cell. 2018. PMID: 28916968 Free PMC article. Review.
-
Optical tweezers experiments resolve distinct modes of DNA-protein binding.Biopolymers. 2009 Apr;91(4):265-82. doi: 10.1002/bip.21123. Biopolymers. 2009. PMID: 19173290 Free PMC article. Review.
Cited by
-
Structure and Functions of HMGB2 Protein.Int J Mol Sci. 2023 May 5;24(9):8334. doi: 10.3390/ijms24098334. Int J Mol Sci. 2023. PMID: 37176041 Free PMC article. Review.
-
HMGB1 deforms nucleosomal DNA to generate a dynamic chromatin environment counteracting the effects of linker histone.Sci Adv. 2025 Aug 15;11(33):eads4473. doi: 10.1126/sciadv.ads4473. Epub 2025 Aug 15. Sci Adv. 2025. PMID: 40815652 Free PMC article.
-
HMGB1 restores a dynamic chromatin environment in the presence of linker histone by deforming nucleosomal DNA.bioRxiv [Preprint]. 2024 Aug 24:2024.08.23.609244. doi: 10.1101/2024.08.23.609244. bioRxiv. 2024. Update in: Sci Adv. 2025 Aug 15;11(33):eads4473. doi: 10.1126/sciadv.ads4473. PMID: 39229246 Free PMC article. Updated. Preprint.
-
Single-molecule analysis of RAG-mediated V(D)J DNA cleavage.Proc Natl Acad Sci U S A. 2015 Apr 7;112(14):E1715-23. doi: 10.1073/pnas.1503477112. Epub 2015 Mar 23. Proc Natl Acad Sci U S A. 2015. PMID: 25831509 Free PMC article.
-
BAP1 forms a trimer with HMGB1 and HDAC1 that modulates gene × environment interaction with asbestos.Proc Natl Acad Sci U S A. 2021 Nov 30;118(48):e2111946118. doi: 10.1073/pnas.2111946118. Proc Natl Acad Sci U S A. 2021. PMID: 34815344 Free PMC article.
References
-
- Bustin M. Revised nomenclature for high mobility group (HMG) chromosomal proteins. Trends Biochem Sci. 2001;26:152–3. - PubMed
-
- Bustin M, Reeves R. High-mobility-group chromosomal proteins: architectural components that facilitate chromatin function. Prog Nucleic Acid Res Mol Biol. 1996;54:35–100. - PubMed
-
- Johns E, editor. HMG Chromosomal Proteins. Academic Press; New York: 1982.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

