Knotted and topologically complex proteins as models for studying folding and stability
- PMID: 17967433
- PMCID: PMC2179896
- DOI: 10.1016/j.cbpa.2007.10.002
Knotted and topologically complex proteins as models for studying folding and stability
Abstract
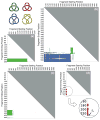
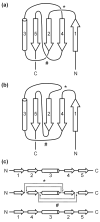
Among proteins of known three-dimensional structure, only a few possess complex topological features such as knotted or interlinked (catenated) protein backbones. Such unusual proteins offer potentially unique insights into folding pathways and stabilization mechanisms. They also present special challenges for both theorists and computational scientists interested in understanding and predicting protein-folding behavior. Here, we review complex topological features in proteins with a focus on recent progress on the identification and characterization of knotted and interlinked protein systems. Also, an approach is described for designing an expanded set of knotted proteins.
Figures

Similar articles
-
Elucidation of folding pathways of knotted proteins.Methods Enzymol. 2022;675:275-297. doi: 10.1016/bs.mie.2022.07.012. Epub 2022 Aug 19. Methods Enzymol. 2022. PMID: 36220273 Review.
-
Tying a true topological protein knot by cyclization.Biochem Biophys Res Commun. 2024 Feb 12;696:149470. doi: 10.1016/j.bbrc.2024.149470. Epub 2024 Jan 12. Biochem Biophys Res Commun. 2024. PMID: 38244314
-
How does a knotted protein fold?FEBS J. 2009 Jan;276(2):365-75. doi: 10.1111/j.1742-4658.2008.06801.x. Epub 2008 Dec 10. FEBS J. 2009. PMID: 19077162 Review.
-
Structures and folding pathways of topologically knotted proteins.J Phys Condens Matter. 2011 Jan 26;23(3):033101. doi: 10.1088/0953-8984/23/3/033101. Epub 2010 Dec 10. J Phys Condens Matter. 2011. PMID: 21406854 Review.
-
Topological descriptions of protein folding.Proc Natl Acad Sci U S A. 2019 May 7;116(19):9360-9369. doi: 10.1073/pnas.1808312116. Epub 2019 Apr 18. Proc Natl Acad Sci U S A. 2019. PMID: 31000594 Free PMC article.
Cited by
-
Hysteresis as a Marker for Complex, Overlapping Landscapes in Proteins.J Phys Chem Lett. 2013 Jan 3;4(1):180-188. doi: 10.1021/jz301893w. Epub 2012 Dec 18. J Phys Chem Lett. 2013. PMID: 23525263 Free PMC article.
-
Mechanically untying a protein slipknot: multiple pathways revealed by force spectroscopy and steered molecular dynamics simulations.J Am Chem Soc. 2012 Jun 27;134(25):10428-35. doi: 10.1021/ja3003205. Epub 2012 Jun 15. J Am Chem Soc. 2012. PMID: 22626004 Free PMC article.
-
Structure and folding of a designed knotted protein.Proc Natl Acad Sci U S A. 2010 Nov 30;107(48):20732-7. doi: 10.1073/pnas.1007602107. Epub 2010 Nov 10. Proc Natl Acad Sci U S A. 2010. PMID: 21068371 Free PMC article.
-
Knotted proteins: A tangled tale of Structural Biology.Comput Struct Biotechnol J. 2015 Aug 19;13:459-68. doi: 10.1016/j.csbj.2015.08.003. eCollection 2015. Comput Struct Biotechnol J. 2015. PMID: 26380658 Free PMC article. Review.
-
The role of non-native interactions in the folding of knotted proteins.PLoS Comput Biol. 2012;8(6):e1002504. doi: 10.1371/journal.pcbi.1002504. Epub 2012 Jun 14. PLoS Comput Biol. 2012. PMID: 22719235 Free PMC article.
References
-
- Plaxco KW, Simons KT, Baker D. Contact order, transition state placement and the refolding rates of single domain proteins. J Mol Biol. 1998;277:985–994. - PubMed
-
- Bryngelson JD, Onuchic JN, Socci ND, Wolynes PG. Funnels, pathways, and the energy landscape of protein folding: a synthesis. Proteins. 1995;21:167–195. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

