Inhibition of CYP2B4 by 2-ethynylnaphthalene: evidence for the co-binding of substrate and inhibitor within the active site
- PMID: 17967439
- PMCID: PMC2121586
- DOI: 10.1016/j.abb.2007.07.032
Inhibition of CYP2B4 by 2-ethynylnaphthalene: evidence for the co-binding of substrate and inhibitor within the active site
Abstract
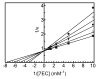
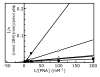
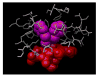
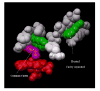
2-ethynylnaphthalene (2EN) is an effective mechanism-based inhibitor of CYP2B4. There are two inhibitory components: (1) irreversible inactivation of CYP2B4 (a typical time-dependent inactivation), and (2) a reversible component. The reversible component was unusual in that the degree of inhibition was not simply a characteristic of the enzyme-inhibitor interaction, but dependent on the size of the substrate molecule used to monitor residual activity. The effect of 2EN on the metabolism of seven CYP2B4 substrates showed that it was not an effective reversible inhibitor of substrates containing a single aromatic ring; substrates with two fused rings were competitively inhibited by 2EN; and larger substrates were non-competitively inhibited. Energy-based docking studies demonstrated that, with increasing substrate size, the energy of 2EN and substrate co-binding in the active site became unfavorable precisely at the point where 2EN became a competitive inhibitor. Hierarchical docking revealed potential allosteric inhibition sites separate from the substrate binding site.
Figures








Similar articles
-
Inhibition of CYP2B4 by the mechanism-based inhibitor 2-ethynylnaphthalene: inhibitory potential of 2EN is dependent on the size of the substrate.Arch Biochem Biophys. 2007 Jun 1;462(1):28-37. doi: 10.1016/j.abb.2007.03.023. Epub 2007 Apr 9. Arch Biochem Biophys. 2007. PMID: 17470357 Free PMC article.
-
Mapping of cytochrome P450 2B4 substrate binding sites by photolabile probe 3-azidiamantane: identification of putative substrate access regions.Arch Biochem Biophys. 2007 Dec 1;468(1):82-91. doi: 10.1016/j.abb.2007.09.017. Epub 2007 Sep 29. Arch Biochem Biophys. 2007. PMID: 17961497
-
Identification of selective mechanism-based inactivators of cytochromes P-450 2B4 and 2B5, and determination of the molecular basis for differential susceptibility.J Pharmacol Exp Ther. 1999 Jul;290(1):445-51. J Pharmacol Exp Ther. 1999. PMID: 10381811
-
Structure-function analysis of cytochromes P450 2B.Biochim Biophys Acta. 2007 Mar;1770(3):402-12. doi: 10.1016/j.bbagen.2006.07.006. Epub 2006 Jul 22. Biochim Biophys Acta. 2007. PMID: 16935426 Review.
-
Differential behavior of the sub-sites of cytochrome 450 active site in binding of substrates, and products (implications for coupling/uncoupling).Biochim Biophys Acta. 2007 Mar;1770(3):360-75. doi: 10.1016/j.bbagen.2006.09.018. Epub 2006 Oct 5. Biochim Biophys Acta. 2007. PMID: 17134838 Review.
Cited by
-
Structure-Function Studies of Naphthalene, Phenanthrene, Biphenyl, and Their Derivatives in Interaction with and Oxidation by Cytochromes P450 2A13 and 2A6.Chem Res Toxicol. 2016 Jun 20;29(6):1029-40. doi: 10.1021/acs.chemrestox.6b00083. Epub 2016 May 12. Chem Res Toxicol. 2016. PMID: 27137136 Free PMC article.
-
Mechanism-based inactivation of cytochrome P450 2C9 by tienilic acid and (+/-)-suprofen: a comparison of kinetics and probe substrate selection.Drug Metab Dispos. 2009 Jan;37(1):59-65. doi: 10.1124/dmd.108.023358. Epub 2008 Oct 6. Drug Metab Dispos. 2009. PMID: 18838506 Free PMC article.
References
-
- White RE, Coon MJ. Annu Rev Biochem. 1980;49:315–356. - PubMed
-
- Korzekwa KR, Krishnamachary N, Shou M, Ogai A, Parise RA, Rettie AE, Gonzalez FJ, Tracy TS. Biochem. 1998;37:4137–4147. - PubMed
-
- Shou M, Dai R, Cui D, Korzekwa KR, Baillie TA, Rushmore TH. J Biol Chem. 2001;276:2256–2262. - PubMed
-
- Hutzler JM, Wienkers LC, Wahlstrom JL, Carlson TJ, Tracy TS. Arch Biochem Biophys. 2003;410:16–24. - PubMed
-
- Ueng YF, Kuwabara T, Chun YJ, Guengerich FP. Biochem. 1997;36:370–381. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

