Identification and characterization of nucleotide-binding site-leucine-rich repeat genes in the model plant Medicago truncatula
- PMID: 17981990
- PMCID: PMC2230567
- DOI: 10.1104/pp.107.104588
Identification and characterization of nucleotide-binding site-leucine-rich repeat genes in the model plant Medicago truncatula
Abstract
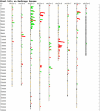
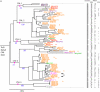
The nucleotide-binding site (NBS)-Leucine-rich repeat (LRR) gene family accounts for the largest number of known disease resistance genes, and is one of the largest gene families in plant genomes. We have identified 333 nonredundant NBS-LRRs in the current Medicago truncatula draft genome (Mt1.0), likely representing 400 to 500 NBS-LRRs in the full genome, or roughly 3 times the number present in Arabidopsis (Arabidopsis thaliana). Although many characteristics of the gene family are similar to those described on other plant genomes, several evolutionary features are particularly pronounced in M. truncatula, including a high degree of clustering, evidence of significant numbers of ectopic translocations from clusters to other parts of the genome, a small number of more evolutionarily stable NBS-LRRs, and numerous truncations and fusions leading to novel domain compositions. The gene family clearly has had a large impact on the structure of the genome, both through ectopic translocations (potentially, a means of seeding new NBS-LRR clusters), and through two extraordinarily large superclusters. Chromosome 6 encodes approximately 34% of all TIR-NBS-LRRs, while chromosome 3 encodes approximately 40% of all coiled-coil-NBS-LRRs. Almost all atypical domain combinations are in the TIR-NBS-LRR subfamily, with many occurring within one genomic cluster. This analysis shows the gene family not only is important functionally and agronomically, but also plays a structural role in the genome.
Figures


References
-
- Axtell MJ, Staskawicz BJ (2003) Initiation of RPS2-specified disease resistance in Arabidopsis is coupled to the AvrRpt2-directed elimination of RIN4. Cell 112 369–377 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

