Synthesis-dependent strand annealing in meiosis
- PMID: 17988174
- PMCID: PMC2062477
- DOI: 10.1371/journal.pbio.0050299
Synthesis-dependent strand annealing in meiosis
Abstract
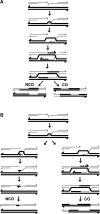
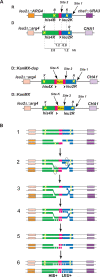
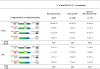
Recent studies led to the proposal that meiotic gene conversion can result after transient engagement of the donor chromatid and subsequent DNA synthesis-dependent strand annealing (SDSA). Double Holliday junction (dHJ) intermediates were previously proposed to form both reciprocal crossover recombinants (COs) and noncrossover recombinants (NCOs); however, dHJs are now thought to give rise mainly to COs, with SDSA forming most or all NCOs. To test this model in Saccharomyces cerevisiae, we constructed a random spore system in which it is possible to identify a subset of NCO recombinants that can readily be accounted for by SDSA, but not by dHJ-mediated recombination. The diagnostic class of recombinants is one in which two markers on opposite sides of a double-strand break site are converted, without conversion of an intervening heterologous insertion located on the donor chromatid. This diagnostic class represents 26% of selected NCO recombinants. Tetrad analysis using the same markers provided additional evidence that SDSA is a major pathway for NCO gene conversion in meiosis.
Conflict of interest statement
Figures

Similar articles
-
Repeated strand invasion and extensive branch migration are hallmarks of meiotic recombination.Mol Cell. 2021 Oct 21;81(20):4258-4270.e4. doi: 10.1016/j.molcel.2021.08.003. Epub 2021 Aug 27. Mol Cell. 2021. PMID: 34453891 Free PMC article.
-
Genome-wide analysis of heteroduplex DNA in mismatch repair-deficient yeast cells reveals novel properties of meiotic recombination pathways.PLoS Genet. 2011 Sep;7(9):e1002305. doi: 10.1371/journal.pgen.1002305. Epub 2011 Sep 29. PLoS Genet. 2011. PMID: 21980306 Free PMC article.
-
Meiotic versus mitotic recombination: two different routes for double-strand break repair: the different functions of meiotic versus mitotic DSB repair are reflected in different pathway usage and different outcomes.Bioessays. 2010 Dec;32(12):1058-66. doi: 10.1002/bies.201000087. Epub 2010 Oct 21. Bioessays. 2010. PMID: 20967781 Free PMC article. Review.
-
Double Holliday junctions are intermediates of DNA break repair.Nature. 2010 Apr 8;464(7290):937-41. doi: 10.1038/nature08868. Epub 2010 Mar 28. Nature. 2010. PMID: 20348905 Free PMC article.
-
Early decision; meiotic crossover interference prior to stable strand exchange and synapsis.Cell. 2004 Apr 2;117(1):9-15. doi: 10.1016/s0092-8674(04)00297-1. Cell. 2004. PMID: 15066278 Review.
Cited by
-
The Role of Chromatid Interference in Determining Meiotic Crossover Patterns.Front Plant Sci. 2021 Mar 9;12:656691. doi: 10.3389/fpls.2021.656691. eCollection 2021. Front Plant Sci. 2021. PMID: 33767725 Free PMC article.
-
Nucleosome occupancy landscape and dynamics at mouse recombination hotspots.EMBO Rep. 2010 Jul;11(7):555-60. doi: 10.1038/embor.2010.79. Epub 2010 May 28. EMBO Rep. 2010. PMID: 20508641 Free PMC article.
-
Meiocyte-Specific and AtSPO11-1-Dependent Small RNAs and Their Association with Meiotic Gene Expression and Recombination.Plant Cell. 2019 Feb;31(2):444-464. doi: 10.1105/tpc.18.00511. Epub 2019 Jan 23. Plant Cell. 2019. PMID: 30674694 Free PMC article.
-
The DNA helicase FANCJ (BRIP1) functions in Double Strand Break repair processing, but not crossover formation during Prophase I of meiosis in male mice.bioRxiv [Preprint]. 2023 Oct 8:2023.10.06.561296. doi: 10.1101/2023.10.06.561296. bioRxiv. 2023. Update in: PLoS Genet. 2024 Feb 20;20(2):e1011175. doi: 10.1371/journal.pgen.1011175. PMID: 37873301 Free PMC article. Updated. Preprint.
-
Mouse tetrad analysis provides insights into recombination mechanisms and hotspot evolutionary dynamics.Nat Genet. 2014 Oct;46(10):1072-80. doi: 10.1038/ng.3068. Epub 2014 Aug 24. Nat Genet. 2014. PMID: 25151354 Free PMC article.
References
-
- Moens PB, Kolas NK, Tarsounas M, Marcon E, Cohen PE, et al. The time course and chromosomal localization of recombination-related proteins at meiosis in the mouse are compatible with models that can resolve the early DNA-DNA interactions without reciprocal recombination. J Cell Sci. 2002;115:1611–1622. - PubMed
-
- Holliday R. Mechanism for gene conversion in fungi. Genetical Res. 1964;5:282–304. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

