WormBase 2007
- PMID: 17991679
- PMCID: PMC2238927
- DOI: 10.1093/nar/gkm975
WormBase 2007
Abstract
WormBase (www.wormbase.org) is the major publicly available database of information about Caenorhabditis elegans, an important system for basic biological and biomedical research. Derived from the initial ACeDB database of C. elegans genetic and sequence information, WormBase now includes the genomic, anatomical and functional information about C. elegans, other Caenorhabditis species and other nematodes. As such, it is a crucial resource not only for C. elegans biologists but the larger biomedical and bioinformatics communities. Coverage of core areas of C. elegans biology will allow the biomedical community to make full use of the results of intensive molecular genetic analysis and functional genomic studies of this organism. Improved search and display tools, wider cross-species comparisons and extended ontologies are some of the features that will help scientists extend their research and take advantage of other nematode species genome sequences.
Figures


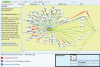
References
-
- TIGR. Brugia Malayi Genome Sequencing Project. 2005. [cited; Available from: http://www.tigr.org/tdb/e2k1/bma1/.
-
- BioMart. [cited; Available from: http://www.biomart.org/].
-
- Matera AG, Terns RM, Terns MP. Non-coding RNAs: lessons from the small nuclear and small nucleolar RNAs. Nat. Rev. Mol. Cell Biol. 2007;8:209–220. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

