Expression of the hepatic specific V1 messenger ribonucleic acid of the human growth hormone receptor gene is regulated by hepatic nuclear factor (HNF)-4alpha2 and HNF-4alpha8
- PMID: 17991764
- PMCID: PMC5419645
- DOI: 10.1210/me.2007-0387
Expression of the hepatic specific V1 messenger ribonucleic acid of the human growth hormone receptor gene is regulated by hepatic nuclear factor (HNF)-4alpha2 and HNF-4alpha8
Abstract
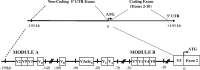
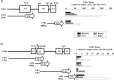
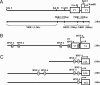
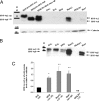
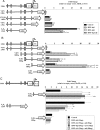
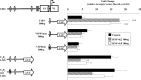
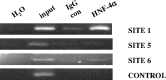
Human (h) GH plays an essential role in growth and metabolism, and its effectiveness is modulated by the availability of its specific receptor [hGH receptor (hGHR)] on target cells. The hGHR gene has a complex 5'-regulatory region containing multiple first exons. Seven are clustered within two small regions: V2,V3,V9 (module A) and V1,V4,V7,V8 (module B). Module A-derived mRNAs are ubiquitously expressed whereas those from module B are only found in postnatal liver, suggesting developmental- and liver-specific regulation of module B hGHR gene expression. To characterize the elements regulating module B activity, we studied a 1.8-kb promoter of the highest expressing exon in liver, V1. This promoter was repressed in transfection assays; however, either 5'- or 3'-deletions relieved this, suggesting the presence of multiple negative regulatory elements. Six putative hepatic nuclear factor 4 (HNF-4) response elements were identified. We determined that HNF-4alpha is developmentally regulated in the human liver: HNF-4alpha2 and HNF-4alpha8 are expressed in fetal hepatocytes but only HNF-4alpha2 is expressed in postnatal liver. Transient transfection assays demonstrated that HNF-4alpha2 and HNF-4alpha8 have a similar dual effect on V1 transcription: activation via site 1 in the proximal promoter and repression through site 6, approximately 1.7 kb upstream. EMSA/electrophoretic mobility supershift assays and chromatin immunoprecipitation analyses confirmed these two sites are bound by HNF-4alpha. Based on these data, we speculate there are multiple regions working together to repress the expression of V1 hGHR transcripts in tissues other than the normal postnatal liver, and that HNF-4alpha is a good candidate for regulating V1 hGHR expression in the human hepatocyte.
Figures



Similar articles
-
Transcription of the human microsomal epoxide hydrolase gene (EPHX1) is regulated by an HNF-4α/CAR/RXR/PSF complex.Biochim Biophys Acta. 2013 Oct;1829(10):1000-9. doi: 10.1016/j.bbagrm.2013.05.003. Epub 2013 May 26. Biochim Biophys Acta. 2013. PMID: 23714182
-
Alu elements in human growth hormone receptor gene 5' untranslated region exons.J Mol Endocrinol. 2001 Dec;27(3):357-66. doi: 10.1677/jme.0.0270357. J Mol Endocrinol. 2001. PMID: 11719288
-
Organization and evolution of the human growth hormone receptor gene 5'-flanking region.Endocrinology. 2001 May;142(5):1923-34. doi: 10.1210/endo.142.5.8170. Endocrinology. 2001. PMID: 11316758
-
Characterization of growth hormone receptor messenger ribonucleic acid variants in human adipocytes.J Clin Endocrinol Metab. 2006 May;91(5):1901-8. doi: 10.1210/jc.2005-1802. Epub 2006 Feb 21. J Clin Endocrinol Metab. 2006. PMID: 16492697
-
Modulation of hepatocyte nuclear factor-4alpha function by the peroxisome-proliferator-activated receptor-gamma co-activator-1alpha in the acute-phase response.Biochem J. 2008 Oct 15;415(2):289-96. doi: 10.1042/BJ20080355. Biochem J. 2008. PMID: 18510493 Free PMC article.
Cited by
-
Genomic structure and characterization of growth hormone receptors from golden pompano Trachinotus ovatus and their expression regulation by feed types.Fish Physiol Biochem. 2019 Dec;45(6):1845-1865. doi: 10.1007/s10695-019-00682-x. Epub 2019 Jul 18. Fish Physiol Biochem. 2019. PMID: 31321605
-
Structure and activity of the human growth hormone receptor (hGHR) gene V2 promoter.Mol Endocrinol. 2009 Mar;23(3):360-72. doi: 10.1210/me.2008-0188. Epub 2008 Dec 30. Mol Endocrinol. 2009. PMID: 19116246 Free PMC article.
-
Regulation of human growth hormone receptor expression by microRNAs.Mol Endocrinol. 2014 Sep;28(9):1448-59. doi: 10.1210/me.2014-1183. Epub 2014 Jul 29. Mol Endocrinol. 2014. PMID: 25073105 Free PMC article.
-
In vivo targeting of the growth hormone receptor (GHR) Box1 sequence demonstrates that the GHR does not signal exclusively through JAK2.Mol Endocrinol. 2010 Jan;24(1):204-17. doi: 10.1210/me.2009-0233. Epub 2009 Nov 2. Mol Endocrinol. 2010. PMID: 19884384 Free PMC article.
References
-
- Veldhuis JD, Roemmich JN, Richmond EJ, Rogol AD, Lovejoy JC, Sheffield-Moore M, Mauras N, Bowers CY 2005. Endocrine control of body composition in infancy, childhood, and puberty. Endocr Rev 26:114–146 - PubMed
-
- Gent J, Van Den EM, van Kerkhof P, Strous GJ 2003. Dimerization and signal transduction of the growth hormone receptor. Mol Endocrinol 17:967–975 - PubMed
-
- Leung DW, Spencer SA, Cachianes G, Hammonds RG, Collins C, Henzel WJ, Barnard R, Waters MJ, Wood WI 1987. Growth hormone receptor and serum binding protein: purification, cloning and expression. Nature 330:537–543 - PubMed
-
- Goodyer CG, Zogopoulos G, Schwartzbauer G, Zheng H, Hendy GN, Menon RK 2001. Organization and evolution of the human growth hormone receptor gene 5′ flanking region. Endocrinology 142:1923–1934 - PubMed
-
- Pekhletsky RI, Chernov BK, Rubtsov PM 1992. Variants of the 5′-untranslated sequence of human growth hormone receptor mRNA. Mol Cell Endocrinol 90:103–109 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

