Discovery of protein interaction networks shared by diseases
- PMID: 17992746
- PMCID: PMC2886192
Discovery of protein interaction networks shared by diseases
Abstract
The study of protein-protein interactions is essential to define the molecular networks that contribute to maintain homeostasis of an organism's body functions. Disruptions in protein interaction networks have been shown to result in diseases in both humans and animals. Monogenic diseases disrupting biochemical pathways such as hereditary coagulopathies (e.g. hemophilia), provided a deep insight in the biochemical pathways of acquired coagulopathies of complex diseases. Indeed, a variety of complex liver diseases can lead to decreased synthesis of the same set of coagulation factors as in hemophilia. Similarly, more complex diseases such as different cancers have been shown to result from malfunctions of common proteins pathways. In order to discover, in high throughput, the molecular underpinnings of poorly characterized diseases, we present a statistical method to identify shared protein interaction network(s) between diseases. Integrating (i) a protein interaction network with (ii) disease to protein relationships derived from mining Gene Ontology annotations and the biomedical literature with natural language understanding (PhenoGO), we identified protein-protein interactions that were associated with pairs of diseases and calculated the statistical significance of the occurrence of interactions in the protein interaction knowledgebase. Significant correlations between diseases and shared protein networks were identified and evaluated in this study, demonstrating the high precision of the approach and correct non-trivial predictions, signifying the potential for discovery. In conclusion, we demonstrate that the associations between diseases are directly correlated to their underlying protein-protein interaction networks, possibly providing insight into the underlying molecular mechanisms of phenotypes and biological processes disrupted in related diseases.
Figures


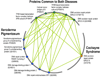
Similar articles
-
The Fritz-Lipmann lecture. DNA repair in human cells. Biochemistry of the hereditary diseases Fanconi's anaemia and Cockayne syndrome.Eur J Biochem. 1987 Jun 1;165(2):235-42. doi: 10.1111/j.1432-1033.1987.tb11433.x. Eur J Biochem. 1987. PMID: 3109898 No abstract available.
-
Pathways for repairing and tolerating the spectrum of oxidative DNA lesions.Cancer Lett. 2012 Dec 31;327(1-2):61-72. doi: 10.1016/j.canlet.2012.02.001. Epub 2012 Feb 19. Cancer Lett. 2012. PMID: 22353689 Free PMC article. Review.
-
Cancer and DNA processing disorders.Br Med Bull. 1994 Jul;50(3):708-17. doi: 10.1093/oxfordjournals.bmb.a072919. Br Med Bull. 1994. PMID: 7987650 Review.
-
Human genetic instability syndromes: single gene defects with increased risk of cancer.Toxicol Lett. 1993 Apr;67(1-3):259-81. doi: 10.1016/0378-4274(93)90061-2. Toxicol Lett. 1993. PMID: 8451764 Review.
-
Diseases with DNA damage-processing defects.Am J Med Sci. 1988 Jan;295(1):40-8. doi: 10.1097/00000441-198801000-00009. Am J Med Sci. 1988. PMID: 3276189 Review.
Cited by
-
Protein-protein interaction networks (PPI) and complex diseases.Gastroenterol Hepatol Bed Bench. 2014 Winter;7(1):17-31. Gastroenterol Hepatol Bed Bench. 2014. PMID: 25436094 Free PMC article. Review.
-
Data driven linear algebraic methods for analysis of molecular pathways: application to disease progression in shock/trauma.J Biomed Inform. 2012 Apr;45(2):372-87. doi: 10.1016/j.jbi.2011.12.002. Epub 2011 Dec 17. J Biomed Inform. 2012. PMID: 22200681 Free PMC article.
-
A vertex similarity-based framework to discover and rank orphan disease-related genes.BMC Syst Biol. 2012;6 Suppl 3(Suppl 3):S8. doi: 10.1186/1752-0509-6-S3-S8. Epub 2012 Dec 17. BMC Syst Biol. 2012. PMID: 23281592 Free PMC article.
-
Evaluation of an Ontology-anchored Natural Language-based Approach for Asserting Multi-scale Biomolecular Networks for Systems Medicine.Summit Transl Bioinform. 2010 Mar 1;2010:6-10. Summit Transl Bioinform. 2010. PMID: 21347135 Free PMC article.
-
A quantitative approach to study indirect effects among disease proteins in the human protein interaction network.BMC Syst Biol. 2010 Jul 29;4:103. doi: 10.1186/1752-0509-4-103. BMC Syst Biol. 2010. PMID: 20670417 Free PMC article.
References
-
- Pinsky L. The polythetic (phenotypic community) system of classifying human malformation syndromes. Birth Defects Orig Artic Ser. 1977;13(3A):13–30. - PubMed
-
- Bertola DR, Kim CA, Albano LM, Scheffer H, Meijer R, van Bokhoven H. Molecular evidence that AEC syndrome and Rapp-Hodgkin syndrome are variable expression of a single genetic disorder. Clin Genet. 2004;66(1):79–80. - PubMed
-
- Sorasio L, Ferrero GB, Garelli E, Brunello G, Martano C, Carando A, et al. Eur J Med Genet. 2006 - PubMed
-
- Zenteno JC, Venegas C, Kofman-Alfaro S. Evidence that AEC syndrome and Bowen--Armstrong syndrome are variable expressions of the same disease. Pediatr Dermatol. 1999;16(2):103–107. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
