Specificity proteins Sp1 and Sp3 interact with the rat GTP cyclohydrolase I proximal promoter to regulate transcription
- PMID: 18004997
- PMCID: PMC2265209
- DOI: 10.1111/j.1471-4159.2007.05054.x
Specificity proteins Sp1 and Sp3 interact with the rat GTP cyclohydrolase I proximal promoter to regulate transcription
Abstract
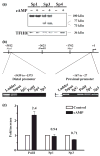
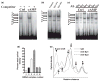
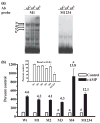
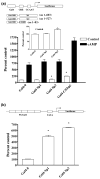
The role of the proximal promoter GC-box in regulating basal and cAMP-dependent GTP Cyclohydrolase I gene transcription was investigated using a variety of cell lines and techniques. These studies show that the GC-box is composed of a triad of cis-elements that in vitro bind specificity proteins Sp1 and Sp3. Sp1 and Sp3 were found associated with the native proximal promoter in PC12 cells but were not recruited to the promoter during cAMP-dependent transcription. Studies using Drosophila SL2 cells showed that Sp3 occupies two sites within the GC-box and enhances transcription when acting alone and synergistically when combined with nuclear factor-Y (NF-Y) and CCAAT/Enhancer-Binding Protein (C/EBP)beta, cognate binding proteins for the adjacent cAMP response element (CRE) and CCAAT-box cAMP response elements. In contrast, Sp1 bound only one site within the GC-box and did not enhance transcription unless combined with NF-Y and C/EBPbeta. Studies in SL2 cells also showed that Sp1 and Sp3 do not co-occupy the GC-box, and accordingly Sp1 competes for Sp3 binding to repress Sp3-dependent transcription. In PC12 cells, complete mutation of the GC-box reduced basal but not cAMP-dependent transcription, resulting in an overall increase in the cAMP response and demonstrating that formation of this enhanceosome does not require Sp1 or Sp3. Experiments in which the GC-box was replaced with a Gal4 element and the promoter challenged with Gal4 fusion proteins support this conclusion and a role for Sp3 in maintaining high levels of basal transcription in PC12 cells. Equivalent amounts of Sp1 and Sp3 were found associated with the native proximal promoter in PC12 and Rat2 cells, which differ 10-fold in basal transcription. Similar levels of methylation of CpG dinucleotides located within the GC-box were also observed in these two cells lines. These results suggest that Sp1 and Sp3 bound to the GC-box might help to preserve an open chromatin configuration at the proximal promoter in cells which constitutively express low levels of GTP Cyclohydrolase I.
Figures



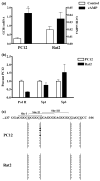
References
-
- Abou-Donia MM, Wilson SP, Zimmerman TP, Nichol CA, Viveros OH. Regulation of guanosine triphosphate cyclohydrolase and tetrahydrobiopterin levels and the role of the cofactor in tyrosine hydroxylation in primary cultures of adrenomedullary chromaffin cells. J Neurochem. 1986;46:1190–1199. - PubMed
-
- Ahlgren R, Suske G, Waterman MR, Lund J. Role of Sp1 in cAMP-dependent transcriptional regulation of the bovine CYP11A gene. J Biol Chem. 1999;274:19422–19428. - PubMed
-
- Anastasiadis PZ, Bezin L, Gordon LJ, Imerman B, Blitz J, Kuhn DM, Levine RA. Vasoactive intestinal peptide induces both tyrosine hydroxylase activity and tetrahydrobiopterin biosynthesis in PC12 cells. Neuroscience. 1998;86:179–189. - PubMed
-
- Bauer M, Suppman S, Meyer M, Hesslinger C, Gasser T, Widner HR, Ueffing M. Glial cell line-derived neurotrophic factor up-regulates GTP-cyclohydrolase I activity and tetrahydrobiopterin levels in primary dopaminergic neurons. J Neurochem. 2002;82:1300–1310. - PubMed
-
- Billon N, Carlisi D, Datto MB, van Grunsven L, Watt A, Wang X-F, Rudkin BB. Cooperation of Sp1 and p300 in the induction of the CDK inhibitor p21WAF1/CIP1 during NGF-mediated neuronal differentiation. Oncogene. 1999;18:2872–2882. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

