Physiological roles of the beta-substituted alanine synthase gene family in Arabidopsis
- PMID: 18024555
- PMCID: PMC2230570
- DOI: 10.1104/pp.107.106831
Physiological roles of the beta-substituted alanine synthase gene family in Arabidopsis
Abstract
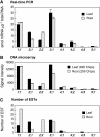
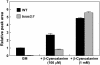
The beta-substituted alanine (Ala) synthase (Bsas) family in the large superfamily of pyridoxal 5'-phosphate-dependent enzymes comprises cysteine (Cys) synthase (CSase) [O-acetyl-serine (thiol) lyase] and beta-cyano-Ala synthase (CASase) in plants. Nine genomic sequences encode putative Bsas proteins in Arabidopsis thaliana. The physiological roles of these Bsas isoforms in vivo were investigated by the characterization of T-DNA insertion mutants. Analyses of gene expression, activities of CSase and CASase, and levels of Cys and glutathione in the bsas mutants indicated that cytosolic Bsas1;1, plastidic Bsas2;1, and mitochondrial Bsas2;2 play major roles in Cys biosynthesis. Cytosolic Bsas1;1 has the most dominant contribution both in leaf and root, and mitochondrial Bsas2;2 plays a significant role in root. Mitochondrial Bsas3;1 is a genuine CASase. Nontargeted metabolome analyses of knockout mutants were carried out by a combination of gas chromatography time-of-flight mass spectrometry and capillary electrophoresis time-of-flight mass spectrometry. The level of gamma-glutamyl-beta-cyano-Ala decreased in the mutant bsas3;1, indicating the crucial role of Bsas3;1 in beta-cyano-Ala metabolism in vivo.
Figures






References
-
- Alonso JM, Stepanova AN, Leisse TJ, Kim CJ, Chen H, Shinn P, Stevenson DK, Zimmerman J, Barajas P, Cheuk R, et al (2003) Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 301 653–657 - PubMed
-
- Anderson ME (1985) Determination of glutathione and glutathione disulfide in biological samples. Methods Enzymol 113 548–555 - PubMed
-
- Ansell M, Lewis FAS (1970) A review of cyanide concentrations found in human organs: a survey of literature concerning cyanide metabolism, “normal,” non-fatal, and fatal body cyanide levels. J Forensic Med 17 148–155 - PubMed
-
- Arabidopsis Genome Initiative (2000) Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408 796–815 - PubMed
-
- Ascano A, Nicholas DJD (1977) Purification and properties of O-acetyl-L-serine sulphydrylase from wheat leaves. Phytochemistry 16 889–893
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

