Combination of two regulatory elements in the Tetrahymena thermophila HSP70-1 gene controls heat shock activation
- PMID: 18055912
- PMCID: PMC2238171
- DOI: 10.1128/EC.00221-07
Combination of two regulatory elements in the Tetrahymena thermophila HSP70-1 gene controls heat shock activation
Abstract
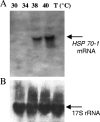
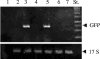
The induction of heat shock genes (HSPs) is thought to be primarily regulated by heat shock transcription factors (HSFs), which bind target sequences on HSP promoters, called heat shock elements (HSEs). In this study, we investigated the 5' untranslated regions of the Tetrahymena thermophila HSP70-1 gene, and we found, in addition to the canonical and divergent HSEs, multiple sets of GATA elements that have not been reported previously in protozoa. By means of in vivo analysis of a green fluorescent protein reporter transgene driven by the HSP70-1 promoter, we demonstrate that HSEs do not represent the minimal regulatory elements for heat shock induction, since the HSP70-1 is tightly regulated by both HSE and GATA elements. Electrophoretic mobility shift assay also showed that HSFs are constitutively bound to the HSEs, whereas GATA elements are engaged only after heat shock. This is the first demonstration by in vivo analysis of functional HSE and GATA elements in protozoa. Furthermore, we provide evidence of a functional link between HSE and GATA elements in the activation of the heat shock response.
Figures



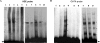
References
-
- Amaral, M. D., L. Galego, and C. Rodrigues-Pousada. 1988. Stress response of Tetrahymena pyriformis to arsenite and heat shock: differences and similarities. Eur. J. Biochem. 171463-470. - PubMed
-
- Amaral, M. D., L. Galero, and C. Rodrigues-Pousada. 1993. Heat-shock-induced protein synthesis is responsible for the switch-off of hsp 70 transcription in Tetrahymena. Biochim. Biophys. Acta 1174133-142. - PubMed
-
- Boorstein, W. R., T. Ziegelhoffer, and E. A. Craig. 1994. Molecular evolution of the HSP70 multigene family. J. Mol. Evol. 381-17. - PubMed
-
- Boye, J., J. Omichinsky, D. Clark, M. Pikaart, and G. Felsenfeld. 1998. Perturbation of nucleosome structure by the erythroid transcription factor GATA-1. J. Mol. Biol. 279529-544. - PubMed
-
- Bradford, M. M. 1976. A rapid and sensitive for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 72248-254. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

