c-myc Repression of TSC2 contributes to control of translation initiation and Myc-induced transformation
- PMID: 18056446
- PMCID: PMC3022657
- DOI: 10.1158/0008-5472.CAN-06-4351
c-myc Repression of TSC2 contributes to control of translation initiation and Myc-induced transformation
Abstract
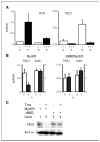
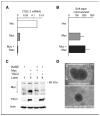
The c-myc oncogene plays a key role in cellular growth control, and translation initiation factors are among the transcriptional targets of Myc. Here, we describe a defect in translation initiation control in myc-null cells due to alterations in the mammalian target of rapamycin (mTOR) pathway. Myc loss increased sensitivity to dominant inhibition of eukaryotic translation initiation factor 4E function. Polysomal profiles of myc(-/-) cells revealed decreased translation initiation rates, which were accompanied by decreased 40S/60S ribosomal subunit ratios. Because the 40S small ribosomal subunit contains the key regulatory ribosomal protein S6 (rpS6), we considered that myc loss might affect expression of components of the mTOR signaling pathway that regulate rpS6 function. Among mTOR signaling components, Myc directly affected transcription of tuberous sclerosis 2 (TSC2), as shown by quantitative mRNA analysis and by Myc binding to its promoter in chromatin immunoprecipitation assays. Importantly, Myc acted as a strong and direct repressor for TSC2 expression because its loss increased TSC2 mRNA in myc-null and in HL60 shRNA experiments, activation of a mycER construct in myc(-/-) cells suppressed TSC2 induction in a myc box II-dependent manner, and mycER activation recruited Myc to the TSC2 promoter. The biological significance of the effect of Myc on TSC2 expression was shown by markedly reduced TSC2 mRNA levels in myc-transformed cells, stimulation of S6 kinase activity in myc-null cells by TSC2 siRNA, and decreased Myc-induced soft agar colony formation following retroviral transduction of TSC2. Together, these findings show that regulation of TSC2 can contribute to the effects of Myc on cell proliferation and neoplastic growth.
Figures




Similar articles
-
The tuberous sclerosis protein TSC2 is not required for the regulation of the mammalian target of rapamycin by amino acids and certain cellular stresses.J Biol Chem. 2005 May 13;280(19):18717-27. doi: 10.1074/jbc.M414499200. Epub 2005 Mar 16. J Biol Chem. 2005. PMID: 15772076
-
RhoE inhibits 4E-BP1 phosphorylation and eIF4E function impairing cap-dependent translation.J Biol Chem. 2009 Dec 18;284(51):35287-96. doi: 10.1074/jbc.M109.050120. J Biol Chem. 2009. PMID: 19850923 Free PMC article.
-
Regulatory effects of mammalian target of rapamycin-activated pathways in type I and II interferon signaling.J Biol Chem. 2007 Jan 19;282(3):1757-68. doi: 10.1074/jbc.M607365200. Epub 2006 Nov 17. J Biol Chem. 2007. PMID: 17114181
-
Growth controls connect: interactions between c-myc and the tuberous sclerosis complex-mTOR pathway.Cell Cycle. 2009 May 1;8(9):1344-51. doi: 10.4161/cc.8.9.8215. Epub 2009 May 18. Cell Cycle. 2009. PMID: 19342893 Free PMC article. Review.
-
Amino acids as regulators of gene expression at the level of mRNA translation.J Nutr. 2003 Jun;133(6 Suppl 1):2046S-2051S. doi: 10.1093/jn/133.6.2046S. J Nutr. 2003. PMID: 12771363 Review.
Cited by
-
Ribosomal protein S15A augments human osteosarcoma cell proliferation in vitro.Cancer Biother Radiopharm. 2014 Dec;29(10):451-6. doi: 10.1089/cbr.2014.1698. Cancer Biother Radiopharm. 2014. Retraction in: Cancer Biother Radiopharm. 2021 Dec;36(10):889. doi: 10.1089/cbr.2014.1698.retract. PMID: 25409460 Free PMC article. Retracted.
-
Exploiting Translation Machinery for Cancer Therapy: Translation Factors as Promising Targets.Int J Mol Sci. 2024 Oct 9;25(19):10835. doi: 10.3390/ijms251910835. Int J Mol Sci. 2024. PMID: 39409166 Free PMC article. Review.
-
MYC-xing it up with PIK3CA mutation and resistance to PI3K inhibitors: summit of two giants in breast cancers.Am J Cancer Res. 2014 Dec 15;5(1):1-19. eCollection 2015. Am J Cancer Res. 2014. PMID: 25628917 Free PMC article. Review.
-
MYC and metabolism on the path to cancer.Semin Cell Dev Biol. 2015 Jul;43:11-21. doi: 10.1016/j.semcdb.2015.08.003. Epub 2015 Aug 12. Semin Cell Dev Biol. 2015. PMID: 26277543 Free PMC article. Review.
-
E2F2 suppresses Myc-induced proliferation and tumorigenesis.Mol Carcinog. 2010 Feb;49(2):152-6. doi: 10.1002/mc.20584. Mol Carcinog. 2010. PMID: 19798698 Free PMC article.
References
-
- Baserga R. The biology of cell reproduction. Cambridge (MA): Harvard University Press; 1985.
-
- Sonenberg N, Hershey JWB, Mathews M, editors. Translational control of gene expression. 2. Cold Spring Harbor (NY): Cold Spring Harbor Laboratory Press; 2000.
-
- Hay N, Sonenberg N. Upstream and downstream of mTOR. Genes Dev. 2004;18:1926–45. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

