Threshold-dominated regulation hides genetic variation in gene expression networks
- PMID: 18062810
- PMCID: PMC2238762
- DOI: 10.1186/1752-0509-1-57
Threshold-dominated regulation hides genetic variation in gene expression networks
Abstract
Background: In dynamical models with feedback and sigmoidal response functions, some or all variables have thresholds around which they regulate themselves or other variables. A mathematical analysis has shown that when the dose-response functions approach binary or on/off responses, any variable with an equilibrium value close to one of its thresholds is very robust to parameter perturbations of a homeostatic state. We denote this threshold robustness. To check the empirical relevance of this phenomenon with response function steepnesses ranging from a near on/off response down to Michaelis-Menten conditions, we have performed a simulation study to investigate the degree of threshold robustness in models for a three-gene system with one downstream gene, using several logical input gates, but excluding models with positive feedback to avoid multistationarity. Varying parameter values representing functional genetic variation, we have analysed the coefficient of variation (CV) of the gene product concentrations in the stable state for the regulating genes in absolute terms and compared to the CV for the unregulating downstream gene. The sigmoidal or binary dose-response functions in these models can be considered as phenomenological models of the aggregated effects on protein or mRNA expression rates of all cellular reactions involved in gene expression.
Results: For all the models, threshold robustness increases with increasing response steepness. The CVs of the regulating genes are significantly smaller than for the unregulating gene, in particular for steep responses. The effect becomes less prominent as steepnesses approach Michaelis-Menten conditions. If the parameter perturbation shifts the equilibrium value too far away from threshold, the gene product is no longer an effective regulator and robustness is lost. Threshold robustness arises when a variable is an active regulator around its threshold, and this function is maintained by the feedback loop that the regulator necessarily takes part in and also is regulated by. In the present study all feedback loops are negative, and our results suggest that threshold robustness is maintained by negative feedback which necessarily exists in the homeostatic state.
Conclusion: Threshold robustness of a variable can be seen as its ability to maintain an active regulation around its threshold in a homeostatic state despite external perturbations. The feedback loop that the system necessarily possesses in this state, ensures that the robust variable is itself regulated and kept close to its threshold. Our results suggest that threshold regulation is a generic phenomenon in feedback-regulated networks with sigmoidal response functions, at least when there is no positive feedback. Threshold robustness in gene regulatory networks illustrates that hidden genetic variation can be explained by systemic properties of the genotype-phenotype map.
Figures

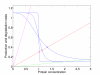

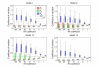
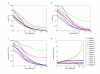
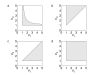
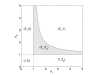
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

