Genomic instability within centromeres of interspecific marsupial hybrids
- PMID: 18073443
- PMCID: PMC2219476
- DOI: 10.1534/genetics.107.082313
Genomic instability within centromeres of interspecific marsupial hybrids
Abstract
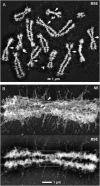
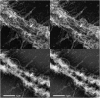
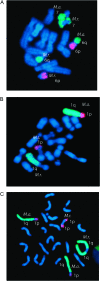
Several lines of evidence suggest that, within a lineage, particular genomic regions are subject to instability that can lead to specific types of chromosome rearrangements important in species incompatibility. Within family Macropodidae (kangaroos, wallabies, bettongs, and potoroos), which exhibit recent and extensive karyotypic evolution, rearrangements involve chiefly the centromere. We propose that centromeres are the primary target for destabilization in cases of genomic instability, such as interspecific hybridization, and participate in the formation of novel chromosome rearrangements. Here we use standard cytological staining, cross-species chromosome painting, DNA probe analyses, and scanning electron microscopy to examine four interspecific macropodid hybrids (Macropus rufogriseus x Macropus agilis). The parental complements share the same centric fusions relative to the presumed macropodid ancestral karyotype, but can be differentiated on the basis of heterochromatic content, M. rufogriseus having larger centromeres with large C-banding positive regions. All hybrids exhibited the same pattern of chromosomal instability and remodeling specifically within the centromeres derived from the maternal (M. rufogriseus) complement. This instability included amplification of a satellite repeat and a transposable element, changes in chromatin structure, and de novo whole-arm rearrangements. We discuss possible reasons and mechanisms for the centromeric instability and remodeling observed in all four macropodid hybrids.
Figures

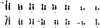

References
-
- Brennecke, J., A. A. Aravin, A. Stark, M. Dus, M. Kellis et al., 2007. Discrete small RNA-generating loci as master regulators of transposon activity in Drosophila. Cell 128: 1089–1103. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

