Localizing frustration in native proteins and protein assemblies
- PMID: 18077414
- PMCID: PMC2148382
- DOI: 10.1073/pnas.0709915104
Localizing frustration in native proteins and protein assemblies
Abstract
We propose a method of quantifying the degree of frustration manifested by spatially local interactions in protein biomolecules. This method of localization smoothly generalizes the global criterion for an energy landscape to be funneled to the native state, which is in keeping with the principle of minimal frustration. A survey of the structural database shows that natural proteins are multiply connected by a web of local interactions that are individually minimally frustrated. In contrast, highly frustrated interactions are found clustered on the surface, often near binding sites. These binding sites become less frustrated upon complex formation.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
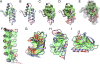




References
-
- Bryngelson JD, Wolynes PG. J Phys Chem. 1989;93:6902–6915.
-
- Clementi C, Maritan A, Banavar JR. Phys Rev Lett. 1998;81:3287–3290.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

