Profiling of UV-induced ATM/ATR signaling pathways
- PMID: 18077418
- PMCID: PMC2148387
- DOI: 10.1073/pnas.0707579104
Profiling of UV-induced ATM/ATR signaling pathways
Abstract
To ensure survival in the face of genomic insult, cells have evolved complex mechanisms to respond to DNA damage, termed the DNA damage checkpoint. The serine/threonine kinases ataxia telangiectasia-mutated (ATM) and ATM and Rad3-related (ATR) activate checkpoint signaling by phosphorylating substrate proteins at SQ/TQ motifs. Although some ATM/ATR substrates (Chk1, p53) have been identified, the lack of a more complete list of substrates limits current understanding of checkpoint pathways. Here, we use immunoaffinity phosphopeptide isolation coupled with mass spectrometry to identify 570 sites phosphorylated in UV-damaged cells, 498 of which are previously undescribed. Semiquantitative analysis yielded 24 known and 192 previously uncharacterized sites differentially phosphorylated upon UV damage, some of which were confirmed by SILAC, Western blotting, and immunoprecipitation/Western blotting. ATR-specific phosphorylation was investigated by using a Seckel syndrome (ATR mutant) cell line. Together, these results provide a rich resource for further deciphering ATM/ATR signaling and the pathways mediating the DNA damage response.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

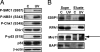

References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

