Understanding the dynamic behavior of genetic regulatory networks by functional decomposition
- PMID: 18079985
- PMCID: PMC2134916
- DOI: 10.2174/138920206778948718
Understanding the dynamic behavior of genetic regulatory networks by functional decomposition
Abstract
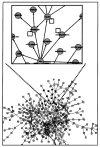
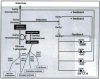
A number of mechanistic and predictive genetic regulatory networks (GRNs) comprising dozens of genes have already been characterized at the level of cis-regulatory interactions. Reconstructions of networks of 100's to 1000's of genes and their interactions are currently underway. Understanding the organizational and functional principles underlying these networks is probably the single greatest challenge facing genomics today. We review the current approaches to deciphering large-scale GRNs and discuss some of their limitations. We then propose a bottom-up approach in which large-scale GRNs are first organized in terms of functionally distinct GRN building blocks of one or a few genes. Biological processes may then be viewed as the outcome of functional interactions among these simple, well-characterized functional building blocks. We describe several putative GRN functional building blocks and show that they can be located within GRNs on the basis of their interaction topology and additional, simple and experimentally testable constraints.
Keywords: Genetic regulatory networks; systems biology; transcriptional regulation; visualization.
Figures




Similar articles
-
MICRAT: a novel algorithm for inferring gene regulatory networks using time series gene expression data.BMC Syst Biol. 2018 Dec 14;12(Suppl 7):115. doi: 10.1186/s12918-018-0635-1. BMC Syst Biol. 2018. PMID: 30547796 Free PMC article.
-
Modeling transcriptional regulatory networks.Bioessays. 2002 Dec;24(12):1118-29. doi: 10.1002/bies.10189. Bioessays. 2002. PMID: 12447977 Review.
-
Periodic synchronization of isolated network elements facilitates simulating and inferring gene regulatory networks including stochastic molecular kinetics.BMC Bioinformatics. 2022 Jan 5;23(1):13. doi: 10.1186/s12859-021-04541-6. BMC Bioinformatics. 2022. PMID: 34986805 Free PMC article.
-
Inference of Gene Regulatory Network Based on Local Bayesian Networks.PLoS Comput Biol. 2016 Aug 1;12(8):e1005024. doi: 10.1371/journal.pcbi.1005024. eCollection 2016 Aug. PLoS Comput Biol. 2016. PMID: 27479082 Free PMC article.
-
Omics approaches to study gene regulatory networks for development in echinoderms.Brief Funct Genomics. 2017 Sep 1;16(5):299-308. doi: 10.1093/bfgp/elx012. Brief Funct Genomics. 2017. PMID: 28957458 Review.
Cited by
-
A bipolar role of the transcription factor ERG for cnidarian germ layer formation and apical domain patterning.Dev Biol. 2017 Oct 15;430(2):346-361. doi: 10.1016/j.ydbio.2017.08.015. Epub 2017 Aug 14. Dev Biol. 2017. PMID: 28818668 Free PMC article.
-
Building developmental gene regulatory networks.Birth Defects Res C Embryo Today. 2009 Jun;87(2):123-30. doi: 10.1002/bdrc.20152. Birth Defects Res C Embryo Today. 2009. PMID: 19530131 Free PMC article. Review.
-
A curated online resource for SOX10 and pigment cell molecular genetic pathways.Database (Oxford). 2010 Oct 25;2010:baq025. doi: 10.1093/database/baq025. Database (Oxford). 2010. PMID: 20974870 Free PMC article.
-
Temperature control of fimbriation circuit switch in uropathogenic Escherichia coli: quantitative analysis via automated model abstraction.PLoS Comput Biol. 2010 Mar 26;6(3):e1000723. doi: 10.1371/journal.pcbi.1000723. PLoS Comput Biol. 2010. PMID: 20361050 Free PMC article.
-
Visualization, documentation, analysis, and communication of large-scale gene regulatory networks.Biochim Biophys Acta. 2009 Apr;1789(4):363-74. doi: 10.1016/j.bbagrm.2008.07.014. Epub 2008 Aug 6. Biochim Biophys Acta. 2009. PMID: 18757046 Free PMC article. Review.
References
-
- Davidson EH, Erwin DH. Gene regulatory networks and the evolution of animal body plans. Science. 2006;311:796–800. - PubMed
-
- Li S, Armstrong CM, Bertin N, Ge H, Milstein S, Boxem M, Vidalain PO, Han JD, Chesneau A, Hao T, Goldberg DS, Li N, Martinez M, Rual JF, Lamesch P, Xu L, Tewari M, Wong SL, Zhang LV, Berriz GF, Jacotot L, Vaglio P, Reboul J, Hirozane–Kishikawa T, Li Q, Gabel HW, Elewa A, Baumgartner B, Rose DJ, Yu H, Bosak S, Sequerra R, Fraser A, Mango SE, Saxton WM, Strome S, Van Den Heuvel S, Piano F, Vandenhaute J, Sardet C, Gerstein M, Doucette-Stamm L, Gunsalus KC, Harper JW, Cusick ME, Roth FP, Hill DE, Vidal M. A map of the interactome network of the metazoan C. elegans. Science. 2004;303:540–3. - PMC - PubMed
-
- Deplancke B, Mukhopadhyay A, Ao W, Elewa AM, Grove CA, Martinez NJ, Sequerra R, Doucette-Stamm L, Reece-Hoyes JS, Hope IA, Tissenbaum HA, Mango SE, Walhout AJ. A gene-centered C. elegans protein-DNA interaction network. Cell. 2006;125:1193–205. - PubMed
-
- Davidson EH. The Regulatory Genome: Gene Regulatory Networks in Development and Evolution. Elsevier; 2006.
-
- Ihmels J, Bergmann S, Barkai N. Defining transcription modules using large-scale gene expression data. Bioinformatics. 2004;20:1993–2003. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous
