CTA4 transcription factor mediates induction of nitrosative stress response in Candida albicans
- PMID: 18083829
- PMCID: PMC2238162
- DOI: 10.1128/EC.00240-07
CTA4 transcription factor mediates induction of nitrosative stress response in Candida albicans
Abstract
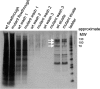
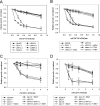
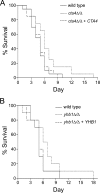
This work has identified regulatory elements in the major fungal pathogen Candida albicans that enable response to nitrosative stress. Nitric oxide (NO) is generated by macrophages of the host immune system and commensal bacteria, and the ability to resist its toxicity is one adaptation that promotes survival of C. albicans inside the human body. Exposing C. albicans to NO induces upregulation of the flavohemoglobin Yhb1p. This protein confers protection by enzymatically converting NO to harmless nitrate, but it is unknown how C. albicans is able to detect NO in its environment and thus initiate this defense only as needed. We analyzed this problem by incrementally mutating the YHB1 regulatory region to identify a nitric oxide-responsive element (NORE) that is required for NO sensitivity. Five transcription factor candidates of the Zn(II)2-Cys6 family were then isolated from crude whole-cell extracts by using magnetic beads coated with this DNA element. Of the five, only deletion of the CTA4 gene prevented induction of YHB1 transcription during nitrosative stress and caused growth sensitivity to the NO donor dipropylenetriamine NONOate; Cta4p associates in vivo with NORE DNA from the YHB1 regulatory region. Deletion of CTA4 caused a small but significant decrease in virulence. A CTA4-dependent putative sulfite transporter encoded by SSU1 is also implicated in NO response, but C. albicans ssu1 mutants were not sensitive to NO, in contrast to findings in Saccharomyces cerevisiae. Cta4p is the first protein found to be necessary for initiating NO response in C. albicans.
Figures



Similar articles
-
Transcriptional response of Candida albicans to nitric oxide and the role of the YHB1 gene in nitrosative stress and virulence.Mol Biol Cell. 2005 Oct;16(10):4814-26. doi: 10.1091/mbc.e05-05-0435. Epub 2005 Jul 19. Mol Biol Cell. 2005. PMID: 16030247 Free PMC article.
-
Adaptation of Candida albicans to Reactive Sulfur Species.Genetics. 2017 May;206(1):151-162. doi: 10.1534/genetics.116.199679. Epub 2017 Feb 24. Genetics. 2017. PMID: 28235888 Free PMC article.
-
In vivo systematic analysis of Candida albicans Zn2-Cys6 transcription factors mutants for mice organ colonization.PLoS One. 2011;6(10):e26962. doi: 10.1371/journal.pone.0026962. Epub 2011 Oct 31. PLoS One. 2011. PMID: 22073120 Free PMC article.
-
Epigenetic Control of Oxidative Stresses by Histone Acetyltransferases in Candida albicans.J Microbiol Biotechnol. 2018 Feb 28;28(2):181-189. doi: 10.4014/jmb.1707.07029. J Microbiol Biotechnol. 2018. PMID: 29169224 Review.
-
From commensal to pathogen: stage- and tissue-specific gene expression of Candida albicans.Curr Opin Microbiol. 2004 Aug;7(4):336-41. doi: 10.1016/j.mib.2004.06.003. Curr Opin Microbiol. 2004. PMID: 15288621 Review.
Cited by
-
Transcriptional Control of Drug Resistance, Virulence and Immune System Evasion in Pathogenic Fungi: A Cross-Species Comparison.Front Cell Infect Microbiol. 2016 Oct 20;6:131. doi: 10.3389/fcimb.2016.00131. eCollection 2016. Front Cell Infect Microbiol. 2016. PMID: 27812511 Free PMC article. Review.
-
The interactions of Candida albicans with gut bacteria: a new strategy to prevent and treat invasive intestinal candidiasis.Gut Pathog. 2023 Jun 27;15(1):30. doi: 10.1186/s13099-023-00559-8. Gut Pathog. 2023. PMID: 37370138 Free PMC article. Review.
-
Interplay of two transcription factors for recruitment of the chromatin remodeling complex modulates fungal nitrosative stress response.Nat Commun. 2021 May 6;12(1):2576. doi: 10.1038/s41467-021-22831-8. Nat Commun. 2021. PMID: 33958593 Free PMC article.
-
R432 is a key residue for the multiple functions of Ndt80p in Candida albicans.Cell Mol Life Sci. 2012 Mar;69(6):1011-23. doi: 10.1007/s00018-011-0849-5. Epub 2011 Oct 16. Cell Mol Life Sci. 2012. PMID: 22002580 Free PMC article.
-
Transcriptional control of hyphal morphogenesis in Candida albicans.FEMS Yeast Res. 2020 Feb 1;20(1):foaa005. doi: 10.1093/femsyr/foaa005. FEMS Yeast Res. 2020. PMID: 31981355 Free PMC article. Review.
References
-
- Akache, B., and B. Turcotte. 2002. New regulators of drug sensitivity in the family of yeast zinc cluster proteins. J. Biol. Chem. 27721254-21260. - PubMed
-
- Aparicio, O., J. V. Geisberg, A. Sekinger, A. Yang, Z. Moqtaderi, and K. Struhl. 2005. Chromatin immunoprecipitation for determining the association of proteins with specific genomic sequences in vivo, p. 1-33. In F. M. Ausubel, R. Brent, R. E. Kingston, D. D. Moore, J. G. Seidman, J. A. Smith, and K. Struhl (ed.), Current protocols in molecular biology. John Wiley and Sons, Inc., New York, NY. - PubMed
-
- Baumgartner, U., B. Hamilton, M. Piskacek, H. Ruis, and H. Rottensteiner. 1999. Functional analysis of the Zn(2)Cys(6) transcription factors Oaf1p and Pip2p. Different roles in fatty acid induction of beta-oxidation in Saccharomyces cerevisiae. J. Biol. Chem. 27422208-22216. - PubMed
-
- Boccara, M., C. E. Mills, J. Zeier, C. Anzi, C. Lamb, R. K. Poole, and M. Delledonne. 2005. Flavohaemoglobin HmpX from Erwinia chrysanthemi confers nitrosative stress tolerance and affects the plant hypersensitive reaction by intercepting nitric oxide produced by the host. Plant J. 43226-237. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

