The genome sequence of the metal-mobilizing, extremely thermoacidophilic archaeon Metallosphaera sedula provides insights into bioleaching-associated metabolism
- PMID: 18083856
- PMCID: PMC2227735
- DOI: 10.1128/AEM.02019-07
The genome sequence of the metal-mobilizing, extremely thermoacidophilic archaeon Metallosphaera sedula provides insights into bioleaching-associated metabolism
Abstract
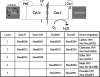
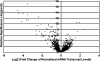
Despite their taxonomic description, not all members of the order Sulfolobales are capable of oxidizing reduced sulfur species, which, in addition to iron oxidation, is a desirable trait of biomining microorganisms. However, the complete genome sequence of the extremely thermoacidophilic archaeon Metallosphaera sedula DSM 5348 (2.2 Mb, approximately 2,300 open reading frames [ORFs]) provides insights into biologically catalyzed metal sulfide oxidation. Comparative genomics was used to identify pathways and proteins involved (directly or indirectly) with bioleaching. As expected, the M. sedula genome contains genes related to autotrophic carbon fixation, metal tolerance, and adhesion. Also, terminal oxidase cluster organization indicates the presence of hybrid quinol-cytochrome oxidase complexes. Comparisons with the mesophilic biomining bacterium Acidithiobacillus ferrooxidans ATCC 23270 indicate that the M. sedula genome encodes at least one putative rusticyanin, involved in iron oxidation, and a putative tetrathionate hydrolase, implicated in sulfur oxidation. The fox gene cluster, involved in iron oxidation in the thermoacidophilic archaeon Sulfolobus metallicus, was also identified. These iron- and sulfur-oxidizing components are missing from genomes of nonleaching members of the Sulfolobales, such as Sulfolobus solfataricus P2 and Sulfolobus acidocaldarius DSM 639. Whole-genome transcriptional response analysis showed that 88 ORFs were up-regulated twofold or more in M. sedula upon addition of ferrous sulfate to yeast extract-based medium; these included genes for components of terminal oxidase clusters predicted to be involved with iron oxidation, as well as genes predicted to be involved with sulfur metabolism. Many hypothetical proteins were also differentially transcribed, indicating that aspects of the iron and sulfur metabolism of M. sedula remain to be identified and characterized.
Figures

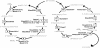


References
-
- Barrett, M. L., I. Harvey, M. Sundararajan, R. Surendran, J. F. Hall, M. J. Ellis, M. A. Hough, R. W. Strange, I. H. Hillier, and S. S. Hasnain. 2006. Atomic resolution crystal structures, EXAFS, and quantum chemical studies of rusticyanin and its two mutants provide insight into its unusual properties. Biochemistry 45:2927-2939. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

