Exploring the molecular etiology of dominant-negative mutations
- PMID: 18083908
- PMCID: PMC2217636
- DOI: 10.1105/tpc.107.055053
Exploring the molecular etiology of dominant-negative mutations
Figures


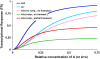
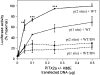
References
-
- Barren, B., and Artemyev, N.O. (2007). Mechanisms of dominant negative G-protein alpha subunits. J. Neurosci. Res. 85 3505–3514. - PubMed
-
- Birchler, J.A., Bhadra, U., Bhadra, M.P., and Auger, D.L. (2001). Dosage-dependent gene regulation in multicellular eukaryotes: implications for dosage compensation, aneuploid syndromes, and quantitative traits. Dev. Biol. 234 275–288. - PubMed
-
- Burz, D.S., and Hanes, S.D. (2001). Isolation of mutations that disrupt cooperative DNA binding by the Drosophila bicoid protein. J. Mol. Biol. 305 219–230. - PubMed
-
- Carey, M. (1998). The enhanceosome and transcriptional synergy. Cell 92 5–8. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

