beta-Catenin is a Nek2 substrate involved in centrosome separation
- PMID: 18086858
- PMCID: PMC2151018
- DOI: 10.1101/gad.1596308
beta-Catenin is a Nek2 substrate involved in centrosome separation
Abstract
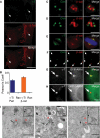
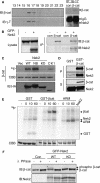
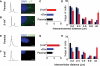
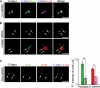
beta-Catenin plays important roles in cell adhesion and gene transcription, and has been shown recently to be essential for the establishment of a bipolar mitotic spindle. Here we show that beta-catenin is a component of interphase centrosomes and that stabilization of beta-catenin, mimicking mutations found in cancers, induces centrosome splitting. Centrosomes are held together by a dynamic linker regulated by Nek2 kinase and its substrates C-Nap1 (centrosomal Nek2-associated protein 1) and Rootletin. We show that beta-catenin binds to and is phosphorylated by Nek2, and is in a complex with Rootletin. In interphase, beta-catenin colocalizes with Rootletin between C-Nap1 puncta at the proximal end of centrioles, and this localization is dependent on C-Nap1 and Rootletin. In mitosis, when Nek2 activity increases, beta-catenin localizes to centrosomes at spindle poles independent of Rootletin. Increased Nek2 activity disrupts the interaction of Rootletin with centrosomes and results in binding of beta-catenin to Rootletin-independent sites on centrosomes, an event that is required for centrosome separation. These results identify beta-catenin as a component of the intercentrosomal linker and define a new function for beta-catenin as a key regulator of mitotic centrosome separation.
Figures


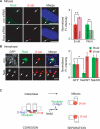
References
-
- Bauer A., Lickert H., Kemler R., Stappert J. Modification of the E-cadherin–catenin complex in mitotic Madin-Darby canine kidney epithelial cells. J. Biol. Chem. 1998;273:28314–28321. - PubMed
-
- Behrens J. The role of the Wnt signalling pathway in colorectal tumorigenesis. Biochem. Soc. Trans. 2005;33:672–675. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous
