Ab initio identification of human microRNAs based on structure motifs
- PMID: 18088431
- PMCID: PMC2238772
- DOI: 10.1186/1471-2105-8-478
Ab initio identification of human microRNAs based on structure motifs
Abstract
Background: MicroRNAs (miRNAs) are short, non-coding RNA molecules that are directly involved in post-transcriptional regulation of gene expression. The mature miRNA sequence binds to more or less specific target sites on the mRNA. Both their small size and sequence specificity make the detection of completely new miRNAs a challenging task. This cannot be based on sequence information alone, but requires structure information about the miRNA precursor. Unlike comparative genomics approaches, ab initio approaches are able to discover species-specific miRNAs without known sequence homology.
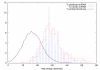
Results: MiRPred is a novel method for ab initio prediction of miRNAs by genome scanning that only relies on (predicted) secondary structure to distinguish miRNA precursors from other similar-sized segments of the human genome. We apply a machine learning technique, called linear genetic programming, to develop special classifier programs which include multiple regular expressions (motifs) matched against the secondary structure sequence. Special attention is paid to scanning issues. The classifiers are trained on fixed-length sequences as these occur when shifting a window in regular steps over a genome region. Various statistical and empirical evidence is collected to validate the correctness of and increase confidence in the predicted structures. Among other things, we propose a new criterion to select miRNA candidates with a higher stability of folding that is based on the number of matching windows around their genome location. An ensemble of 16 motif-based classifiers achieves 99.9 percent specificity with sensitivity remaining on an acceptable high level when requiring all classifiers to agree on a positive decision. A low false positive rate is considered more important than a low false negative rate, when searching larger genome regions for unknown miRNAs. 117 new miRNAs have been predicted close to known miRNAs on human chromosome 19. All candidate structures match the free energy distribution of miRNA precursors which is significantly shifted towards lower free energies. We employed a human EST library and found that around 75 percent of the candidate sequences are likely to be transcribed, with around 35 percent located in introns.
Conclusion: Our motif finding method is at least competitive to state-of-the-art feature-based methods for ab initio miRNA discovery. In doing so, it requires less previous knowledge about miRNA precursor structures while programs and motifs allow a more straightforward interpretation and extraction of the acquired knowledge.
Figures



Similar articles
-
Classification of real and pseudo microRNA precursors using local structure-sequence features and support vector machine.BMC Bioinformatics. 2005 Dec 29;6:310. doi: 10.1186/1471-2105-6-310. BMC Bioinformatics. 2005. PMID: 16381612 Free PMC article.
-
Mirnacle: machine learning with SMOTE and random forest for improving selectivity in pre-miRNA ab initio prediction.BMC Bioinformatics. 2016 Dec 15;17(Suppl 18):474. doi: 10.1186/s12859-016-1343-8. BMC Bioinformatics. 2016. PMID: 28105918 Free PMC article.
-
Predicting microRNA precursors with a generalized Gaussian components based density estimation algorithm.BMC Bioinformatics. 2010 Jan 18;11 Suppl 1(Suppl 1):S52. doi: 10.1186/1471-2105-11-S1-S52. BMC Bioinformatics. 2010. PMID: 20122227 Free PMC article.
-
[Computational approaches to microRNA discovery].Yi Chuan. 2008 Jun;30(6):687-96. doi: 10.3724/sp.j.1005.2008.00687. Yi Chuan. 2008. PMID: 18550489 Review. Chinese.
-
Popular Computational Tools Used for miRNA Prediction and Their Future Development Prospects.Interdiscip Sci. 2020 Dec;12(4):395-413. doi: 10.1007/s12539-020-00387-3. Epub 2020 Sep 21. Interdiscip Sci. 2020. PMID: 32959233 Review.
Cited by
-
Identification of real microRNA precursors with a pseudo structure status composition approach.PLoS One. 2015 Mar 30;10(3):e0121501. doi: 10.1371/journal.pone.0121501. eCollection 2015. PLoS One. 2015. PMID: 25821974 Free PMC article.
-
Computational and experimental identification of mirtrons in Drosophila melanogaster and Caenorhabditis elegans.Genome Res. 2011 Feb;21(2):286-300. doi: 10.1101/gr.113050.110. Epub 2010 Dec 22. Genome Res. 2011. PMID: 21177960 Free PMC article.
-
miRBoost: boosting support vector machines for microRNA precursor classification.RNA. 2015 May;21(5):775-85. doi: 10.1261/rna.043612.113. Epub 2015 Mar 20. RNA. 2015. PMID: 25795417 Free PMC article.
-
Computational Characterization of Exogenous MicroRNAs that Can Be Transferred into Human Circulation.PLoS One. 2015 Nov 3;10(11):e0140587. doi: 10.1371/journal.pone.0140587. eCollection 2015. PLoS One. 2015. Retraction in: PLoS One. 2022 May 9;17(5):e0268437. doi: 10.1371/journal.pone.0268437. PMID: 26528912 Free PMC article. Retracted.
-
MiRPara: a SVM-based software tool for prediction of most probable microRNA coding regions in genome scale sequences.BMC Bioinformatics. 2011 Apr 19;12:107. doi: 10.1186/1471-2105-12-107. BMC Bioinformatics. 2011. PMID: 21504621 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

