Anaerobic metabolism of catechol by the denitrifying bacterium Thauera aromatica--a result of promiscuous enzymes and regulators?
- PMID: 18156265
- PMCID: PMC2258688
- DOI: 10.1128/JB.01221-07
Anaerobic metabolism of catechol by the denitrifying bacterium Thauera aromatica--a result of promiscuous enzymes and regulators?
Abstract
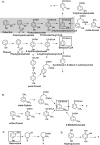
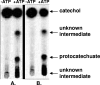
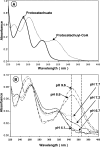
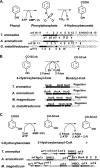
The anaerobic metabolism of catechol (1,2-dihydroxybenzene) was studied in the betaproteobacterium Thauera aromatica that was grown with CO2 as a cosubstrate and nitrate as an electron acceptor. Based on different lines of evidence and on our knowledge of enzymes and genes involved in the anaerobic metabolism of other aromatic substrates, the following pathway is proposed. Catechol is converted to catechylphosphate by phenylphosphate synthase, which is followed by carboxylation by phenylphosphate carboxylase at the para position to the phosphorylated phenolic hydroxyl group. The product, protocatechuate (3,4-dihydroxybenzoate), is converted to its coenzyme A (CoA) thioester by 3-hydroxybenzoate-CoA ligase. Protocatechuyl-CoA is reductively dehydroxylated to 3-hydroxybenzoyl-CoA, possibly by 4-hydroxybenzoyl-CoA reductase. 3-Hydroxybenzoyl-CoA is further metabolized by reduction of the aromatic ring catalyzed by an ATP-driven benzoyl-CoA reductase. Hence, the promiscuity of several enzymes and regulatory proteins may be sufficient to create the catechol pathway that is made up of elements of phenol, 3-hydroxybenzoate, 4-hydroxybenzoate, and benzoate metabolism.
Figures


Similar articles
-
Anaerobic metabolism of 3-hydroxybenzoate by the denitrifying bacterium Thauera aromatica.J Bacteriol. 2001 Feb;183(3):968-79. doi: 10.1128/JB.183.3.968-979.2001. J Bacteriol. 2001. PMID: 11208796 Free PMC article.
-
Phenylphosphate carboxylase: a new C-C lyase involved in anaerobic phenol metabolism in Thauera aromatica.J Bacteriol. 2004 Jul;186(14):4556-67. doi: 10.1128/JB.186.14.4556-4567.2004. J Bacteriol. 2004. PMID: 15231788 Free PMC article.
-
Enzymes involved in the anaerobic degradation of phenol by the sulfate-reducing bacterium Desulfatiglans anilini.BMC Microbiol. 2018 Aug 29;18(1):93. doi: 10.1186/s12866-018-1238-0. BMC Microbiol. 2018. PMID: 30157755 Free PMC article.
-
Unusual reactions involved in anaerobic metabolism of phenolic compounds.Biol Chem. 2005 Oct;386(10):989-97. doi: 10.1515/BC.2005.115. Biol Chem. 2005. PMID: 16218871 Review.
-
Key enzymes in the anaerobic aromatic metabolism catalysing Birch-like reductions.Biochim Biophys Acta. 2005 Feb 25;1707(1):34-50. doi: 10.1016/j.bbabio.2004.01.009. Biochim Biophys Acta. 2005. PMID: 15721605 Review.
Cited by
-
Anaerobic catabolism of aromatic compounds: a genetic and genomic view.Microbiol Mol Biol Rev. 2009 Mar;73(1):71-133. doi: 10.1128/MMBR.00021-08. Microbiol Mol Biol Rev. 2009. PMID: 19258534 Free PMC article. Review.
-
Non-Oxidative Enzymatic (De)Carboxylation of (Hetero)Aromatics and Acrylic Acid Derivatives.Adv Synth Catal. 2019 Jun 6;361(11):2402-2420. doi: 10.1002/adsc.201900275. Epub 2019 May 17. Adv Synth Catal. 2019. PMID: 31379472 Free PMC article. Review.
-
Benzoyl coenzyme a pathway-mediated metabolism of meta-hydroxy-aromatic acids in Rhodopseudomonas palustris.J Bacteriol. 2013 Sep;195(18):4112-20. doi: 10.1128/JB.00634-13. Epub 2013 Jul 12. J Bacteriol. 2013. PMID: 23852864 Free PMC article.
-
Microbial diversity, community composition and metabolic potential in hydrocarbon contaminated oily sludge: prospects for in situ bioremediation.Environ Sci Pollut Res Int. 2014 Jun;21(12):7369-89. doi: 10.1007/s11356-014-2640-2. Epub 2014 Mar 1. Environ Sci Pollut Res Int. 2014. PMID: 24682711
-
Methanogenic degradation of lignin-derived monoaromatic compounds by microbial enrichments from rice paddy field soil.Sci Rep. 2015 Sep 24;5:14295. doi: 10.1038/srep14295. Sci Rep. 2015. PMID: 26399549 Free PMC article.
References
-
- Anders, H. J., A. Kaetzke, P. Kampfer, W. Ludwig, and G. Fuchs. 1995. Taxonomic position of aromatic-degrading denitrifying Pseudomonas strains K 172 and KB 740 and their description as new members of the genera Thauera, as Thauera aromatica sp. nov., and Azoarcus, as Azoarcus evansii sp. nov., respectively, members of the beta subclass of the Proteobacteria. Int. J. Syst. Bacteriol. 45327-333. - PubMed
-
- Biegert, T., U. Altenschmidt, C. Eckerskorn, and G. Fuchs. 1993. Enzymes of anaerobic metabolism of phenolic compounds. 4-Hydroxybenzoate-CoA ligase from a denitrifying Pseudomonas species. Eur. J. Biochem. 213555-561. - PubMed
-
- Bonting, C. F., and G. Fuchs. 1996. Anaerobic metabolism of 2-hydroxybenzoic acid (salicylic acid) by a denitrifying bacterium. Arch. Microbiol. 165402-408. - PubMed
-
- Bradford, M. M. 1976. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 72248-252. - PubMed
-
- Breese, K., and G. Fuchs. 1998. 4-Hydroxybenzoyl-CoA reductase (dehydroxylating) from the denitrifying bacterium Thauera aromatica—prosthetic groups, electron donor and genes of a member of the molybdenum-flavin-iron-sulfur proteins. Eur. J. Biochem. 251916-923. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources

