Anthocyanin inhibits propidium iodide DNA fluorescence in Euphorbia pulcherrima: implications for genome size variation and flow cytometry
- PMID: 18158306
- PMCID: PMC2710214
- DOI: 10.1093/aob/mcm303
Anthocyanin inhibits propidium iodide DNA fluorescence in Euphorbia pulcherrima: implications for genome size variation and flow cytometry
Abstract
Background: Measuring genome size by flow cytometry assumes direct proportionality between nuclear DNA staining and DNA amount. By 1997 it was recognized that secondary metabolites may affect DNA staining, thereby causing inaccuracy. Here experiments are reported with poinsettia (Euphorbia pulcherrima) with green leaves and red bracts rich in phenolics.
Methods: DNA content was estimated as fluorescence of propidium iodide (PI)-stained nuclei of poinsettia and/or pea (Pisum sativum) using flow cytometry. Tissue was chopped, or two tissues co-chopped, in Galbraith buffer alone or with six concentrations of cyanidin-3-rutinoside (a cyanidin-3-rhamnoglucoside contributing to red coloration in poinsettia).
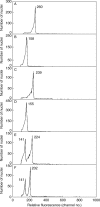
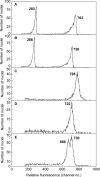
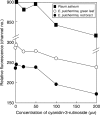
Key results: There were large differences in PI staining (35-70 %) between 2C nuclei from green leaf and red bract tissue in poinsettia. These largely disappeared when pea leaflets were co-chopped with poinsettia tissue as an internal standard. However, smaller (2.8-6.9 %) differences remained, and red bracts gave significantly lower 1C genome size estimates (1.69-1.76 pg) than green leaves (1.81 pg). Chopping pea or poinsettia tissue in buffer with 0-200 microm cyanidin-3-rutinoside showed that the effects of natural inhibitors in red bracts of poinsettia on PI staining were largely reproduced in a dose-dependent way by this anthocyanin.
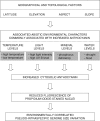
Conclusions: Given their near-ubiquitous distribution, many suspected roles and known affects on DNA staining, anthocyanins are a potent, potential cause of significant error variation in genome size estimations for many plant tissues and taxa. This has important implications of wide practical and theoretical significance. When choosing genome size calibration standards it seems prudent to select materials producing little or no anthocyanin. Reviewing the literature identifies clear examples in which claims of intraspecific variation in genome size are probably artefacts caused by natural variation in anthocyanin levels or correlated with environmental factors known to induce variation in pigmentation.
Figures
References
-
- Arabidopsis Genome Initiative. Analysis of the genome sequence of the flowering plant. Arabidopsis thaliana. Nature. 2000;408:796–815. - PubMed
-
- Arumuganathan KE, Earle ED. Nuclear DNA content of some important plant species. Plant Molecular Biology Reporter. 1991;9:208–219.
-
- Bassi P. Quantitative variations of nuclear DNA during plant development: a critical analysis. Biological Reviews of the Cambridge Philosophical Society. 1990;65:185–225.
-
- Bennett MD. Intraspecific variation in DNA amount and the nucleotypic dimension in plant genetics. In: Freeling M, editor. UCLA Symposium on molecular and cellular biology, Plant genetics. Vol. 35. New York: Alan Liss; 1985. pp. 283–302.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

