Multiple DNA-binding sites in Tetrahymena telomerase
- PMID: 18174223
- PMCID: PMC2275084
- DOI: 10.1093/nar/gkm866
Multiple DNA-binding sites in Tetrahymena telomerase
Abstract
Telomerase is a ribonucleoprotein enzyme that maintains chromosome ends through de novo addition of telomeric DNA. The ability of telomerase to interact with its DNA substrate at sites outside its catalytic centre ('anchor sites') is important for its unique ability to undergo repeat addition processivity. We have developed a direct and quantitative equilibrium primer-binding assay to measure DNA-binding affinities of regions of the catalytic protein subunit of recombinant Tetrahymena telomerase (TERT). There are specific telomeric DNA-binding sites in at least four regions of TERT (the TEN, RBD, RT and C-terminal domains). Together, these sites contribute to specific and high-affinity DNA binding, with a K(d) of approximately 8 nM. Both the K(m) and K(d) increased in a stepwise manner as the primer length was reduced; thus recombinant Tetrahymena telomerase, like the endogenous enzyme, contains multiple anchor sites. The N-terminal TEN domain, which has previously been implicated in DNA binding, shows only low affinity binding. However, there appears to be cooperativity between the TEN and RNA-binding domains. Our data suggest that different DNA-binding sites are used by the enzyme during different stages of the addition cycle.
Figures


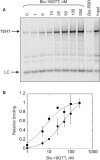




Similar articles
-
Mutation in TERT separates processivity from anchor-site function.Nat Struct Mol Biol. 2008 Aug;15(8):870-2. doi: 10.1038/nsmb.1462. Epub 2008 Jul 20. Nat Struct Mol Biol. 2008. PMID: 18641663 Free PMC article.
-
A conserved motif in Tetrahymena thermophila telomerase reverse transcriptase is proximal to the RNA template and is essential for boundary definition.J Biol Chem. 2013 Jul 26;288(30):22141-9. doi: 10.1074/jbc.M113.452425. Epub 2013 Jun 11. J Biol Chem. 2013. PMID: 23760279 Free PMC article.
-
The architecture of Tetrahymena telomerase holoenzyme.Nature. 2013 Apr 11;496(7444):187-92. doi: 10.1038/nature12062. Epub 2013 Apr 3. Nature. 2013. PMID: 23552895 Free PMC article.
-
xRRM: a new class of RRM found in the telomerase La family protein p65.RNA Biol. 2013 Mar;10(3):353-9. doi: 10.4161/rna.23608. Epub 2013 Jan 17. RNA Biol. 2013. PMID: 23328630 Free PMC article. Review.
-
Progress in structural studies of telomerase.Curr Opin Struct Biol. 2014 Feb;24:115-24. doi: 10.1016/j.sbi.2014.01.008. Epub 2014 Feb 4. Curr Opin Struct Biol. 2014. PMID: 24508601 Free PMC article. Review.
Cited by
-
A self-regulating template in human telomerase.Proc Natl Acad Sci U S A. 2014 Aug 5;111(31):11311-6. doi: 10.1073/pnas.1402531111. Epub 2014 Jun 30. Proc Natl Acad Sci U S A. 2014. PMID: 24982163 Free PMC article.
-
Tracing the path of DNA substrates in active Tetrahymena telomerase holoenzyme complexes: mapping of DNA contact sites in the RNA subunit.Nucleic Acids Res. 2012 Aug;40(15):7430-41. doi: 10.1093/nar/gks416. Epub 2012 May 14. Nucleic Acids Res. 2012. PMID: 22584626 Free PMC article.
-
Ribonucleoprotein multimers and their functions.Crit Rev Biochem Mol Biol. 2010 Oct;45(5):331-50. doi: 10.3109/10409238.2010.496772. Crit Rev Biochem Mol Biol. 2010. PMID: 20572804 Free PMC article. Review.
-
Human telomerase specialization for repeat synthesis by unique handling of primer-template duplex.EMBO J. 2014 Apr 16;33(8):921-35. doi: 10.1002/embj.201387205. Epub 2014 Mar 11. EMBO J. 2014. PMID: 24619002 Free PMC article.
-
Inhibition of yeast telomerase action by the telomeric ssDNA-binding protein, Cdc13p.Nucleic Acids Res. 2009 Feb;37(2):354-67. doi: 10.1093/nar/gkn830. Epub 2008 Nov 29. Nucleic Acids Res. 2009. PMID: 19043074 Free PMC article.
References
-
- McEachern MJ, Krauskopf A, Blackburn EH. Telomeres and their control. Annu. Rev. Genet. 2000;34:331–358. - PubMed
-
- Greider CW, Blackburn EH. Identification of a specific telomere terminal transferase activity in Tetrahymena extracts. Cell. 1985;43:405–413. - PubMed
-
- Shay JW, Bacchetti S. A survey of telomerase activity in human cancer. Eur. J. Cancer. 1997;33:787–791. - PubMed
-
- Hahn WC, Stewart SA, Brooks MW, York SG, Eaton E, Kurachi A, Beijersbergen RL, Knoll JH, Meyerson M, et al. Inhibition of telomerase limits the growth of human cancer cells. Nat. Med. 1999;5:1164–1170. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

