Extending pathways based on gene lists using InterPro domain signatures
- PMID: 18177498
- PMCID: PMC2245903
- DOI: 10.1186/1471-2105-9-3
Extending pathways based on gene lists using InterPro domain signatures
Abstract
Background: High-throughput technologies like functional screens and gene expression analysis produce extended lists of candidate genes. Gene-Set Enrichment Analysis is a commonly used and well established technique to test for the statistically significant over-representation of particular pathways. A shortcoming of this method is however, that most genes that are investigated in the experiments have very sparse functional or pathway annotation and therefore cannot be the target of such an analysis. The approach presented here aims to assign lists of genes with limited annotation to previously described functional gene collections or pathways. This works by comparing InterPro domain signatures of the candidate gene lists with domain signatures of gene sets derived from known classifications, e.g. KEGG pathways.
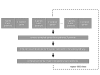
Results: In order to validate our approach, we designed a simulation study. Based on all pathways available in the KEGG database, we create test gene lists by randomly selecting pathway genes, removing these genes from the known pathways and adding variable amounts of noise in the form of genes not annotated to the pathway. We show that we can recover pathway memberships based on the simulated gene lists with high accuracy. We further demonstrate the applicability of our approach on a biological example.
Conclusion: Results based on simulation and data analysis show that domain based pathway enrichment analysis is a very sensitive method to test for enrichment of pathways in sparsely annotated lists of genes. An R based software package domainsignatures, to routinely perform this analysis on the results of high-throughput screening, is available via Bioconductor.
Figures


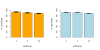
Similar articles
-
How to decide which are the most pertinent overly-represented features during gene set enrichment analysis.BMC Bioinformatics. 2007 Sep 11;8:332. doi: 10.1186/1471-2105-8-332. BMC Bioinformatics. 2007. PMID: 17848190 Free PMC article.
-
SEGS: search for enriched gene sets in microarray data.J Biomed Inform. 2008 Aug;41(4):588-601. doi: 10.1016/j.jbi.2007.12.001. Epub 2007 Dec 15. J Biomed Inform. 2008. PMID: 18234563
-
Reliable gene signatures for microarray classification: assessment of stability and performance.Bioinformatics. 2006 Oct 1;22(19):2356-63. doi: 10.1093/bioinformatics/btl400. Epub 2006 Jul 31. Bioinformatics. 2006. PMID: 16882647
-
Searching for hypothetical proteins: theory and practice based upon original data and literature.Prog Neurobiol. 2005 Sep-Oct;77(1-2):90-127. doi: 10.1016/j.pneurobio.2005.10.001. Epub 2005 Nov 4. Prog Neurobiol. 2005. PMID: 16271823 Review.
-
Stability and aggregation of ranked gene lists.Brief Bioinform. 2009 Sep;10(5):556-68. doi: 10.1093/bib/bbp034. Brief Bioinform. 2009. PMID: 19679825 Review.
Cited by
-
REFINING CELLULAR PATHWAY MODELS USING AN ENSEMBLE OF HETEROGENEOUS DATA SOURCES.Ann Appl Stat. 2018 Sep;12(3):1361-1384. doi: 10.1214/16-aoas915. Epub 2018 Sep 11. Ann Appl Stat. 2018. PMID: 36506698 Free PMC article.
-
PhenoFam-gene set enrichment analysis through protein structural information.BMC Bioinformatics. 2010 May 17;11:254. doi: 10.1186/1471-2105-11-254. BMC Bioinformatics. 2010. PMID: 20478033 Free PMC article.
-
Pathways-based analyses of whole-genome association study data in bipolar disorder reveal genes mediating ion channel activity and synaptic neurotransmission.Hum Genet. 2009 Feb;125(1):63-79. doi: 10.1007/s00439-008-0600-y. Epub 2008 Dec 4. Hum Genet. 2009. PMID: 19052778
-
Successful in vitro expansion and differentiation of cord blood derived CD34+ cells into early endothelial progenitor cells reveals highly differential gene expression.PLoS One. 2011;6(8):e23210. doi: 10.1371/journal.pone.0023210. Epub 2011 Aug 12. PLoS One. 2011. PMID: 21858032 Free PMC article.
-
Leveraging Comparative Genomics to Identify and Functionally Characterize Genes Associated with Sperm Phenotypes in Python bivittatus (Burmese Python).Genet Res Int. 2016;2016:7505268. doi: 10.1155/2016/7505268. Epub 2016 Apr 20. Genet Res Int. 2016. PMID: 27200191 Free PMC article.
References
-
- Al-Shahrour F, Minguez P, Tárraga J, Medina I, Alloza E, Montaner D, Dopazo J. FatiGO +: a functional profiling tool for genomic data. Integration of functional annotation, regulatory motifs and interaction data with microarray experiments. Nucleic Acids Res. 2007;35:W91–W96. doi: 10.1093/nar/gkm260. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

