Epidermal growth factor receptor and protein kinase C signaling to ERK2: spatiotemporal regulation of ERK2 by dual specificity phosphatases
- PMID: 18178562
- PMCID: PMC2268109
- DOI: 10.1074/jbc.M706624200
Epidermal growth factor receptor and protein kinase C signaling to ERK2: spatiotemporal regulation of ERK2 by dual specificity phosphatases
Abstract
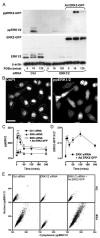
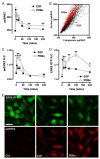
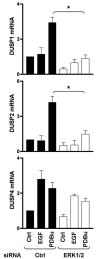
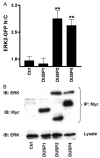
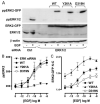
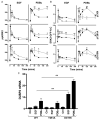
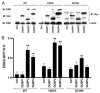
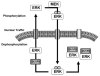
Spatiotemporal aspects of ERK activation are stimulus-specific and dictate cellular consequences. They are dependent upon dual specificity phosphatases (DUSPs) that bind ERK via docking domains and can both inactivate and anchor ERK in cellular compartments. Using high throughput fluorescence microscopy in combination with a system where endogenous ERKs are removed and replaced with wild-type or mutated ERK2-green fluorescent protein (GFP), we show that ERK2 activation responses to epidermal growth factor (EGF) and protein kinase C (PKC) are transient and sustained, respectively. PKC-mediated ERK2 activation is associated with prolonged nuclear localization in the dephosphorylated form, whereas EGF-stimulated ERK2 activation mediates only transient nuclear accumulation. By using short inhibitory RNAs to nuclear inducible DUSP1, -2, or -4 (alone or in combination), we demonstrate that all three of these enzymes contribute to the dephosphorylation of PKC (but not EGF)-activated ERK2 in the nucleus but that they have opposing effects on localization. DUSP2 and -4 inactivate and anchor ERK2, whereas DUSP1 dephosphorylates ERK in the nucleus but allows its traffic back to the cytoplasm. Overexpression of DUSP1, -2, or -4 prevented ERK2 activation, but only DUSP2 and -4 caused ERK2-GFP nuclear accumulation or could be immunoprecipitated with ERK2. Furthermore, protein synthesis inhibition or replacement of wild-type ERK2-GFP with docking domain mutants selectively increased PKC effects on ERK activity and altered ERK2-GFP localization. These mutations also impaired the ability of ERK2-GFP to bind DUSP2 and -4. Together, our data reveal a novel, stimulus-specific, and phosphatase-specific mechanism of ERK2 regulation in the nucleus by DUSP1, -2, and -4.
Figures


Similar articles
-
Spatiotemporal regulation of ERK2 by dual specificity phosphatases.J Biol Chem. 2008 Sep 26;283(39):26612-23. doi: 10.1074/jbc.M801500200. Epub 2008 Jul 23. J Biol Chem. 2008. PMID: 18650424 Free PMC article.
-
Gonadotropin-releasing hormone and protein kinase C signaling to ERK: spatiotemporal regulation of ERK by docking domains and dual-specificity phosphatases.Mol Endocrinol. 2009 Apr;23(4):510-9. doi: 10.1210/me.2008-0333. Epub 2009 Jan 29. Mol Endocrinol. 2009. PMID: 19179479 Free PMC article.
-
Dual specificity phosphatases 10 and 16 are positive regulators of EGF-stimulated ERK activity: indirect regulation of ERK signals by JNK/p38 selective MAPK phosphatases.Cell Signal. 2012 May;24(5):1002-11. doi: 10.1016/j.cellsig.2011.12.021. Epub 2012 Jan 3. Cell Signal. 2012. PMID: 22245064 Free PMC article.
-
Membrane proximal ERK signaling is required for M-calpain activation downstream of epidermal growth factor receptor signaling.J Biol Chem. 2001 Jun 29;276(26):23341-8. doi: 10.1074/jbc.M008847200. Epub 2001 Apr 23. J Biol Chem. 2001. PMID: 11319218
-
Specific inactivation and nuclear anchoring of extracellular signal-regulated kinase 2 by the inducible dual-specificity protein phosphatase DUSP5.Mol Cell Biol. 2005 Mar;25(5):1830-45. doi: 10.1128/MCB.25.5.1830-1845.2005. Mol Cell Biol. 2005. PMID: 15713638 Free PMC article.
Cited by
-
Noncanonical regulation of insulin-mediated ERK activation by phosphoinositide 3-kinase γ.Mol Biol Cell. 2017 Nov 1;28(22):3112-3122. doi: 10.1091/mbc.E16-12-0864. Epub 2017 Sep 6. Mol Biol Cell. 2017. PMID: 28877982 Free PMC article.
-
Receptor type protein tyrosine phosphatase-kappa mediates cross-talk between transforming growth factor-beta and epidermal growth factor receptor signaling pathways in human keratinocytes.Mol Biol Cell. 2010 Jan 1;21(1):29-35. doi: 10.1091/mbc.e09-08-0710. Epub 2009 Oct 28. Mol Biol Cell. 2010. PMID: 19864455 Free PMC article.
-
Spatiotemporal regulation of ERK2 by dual specificity phosphatases.J Biol Chem. 2008 Sep 26;283(39):26612-23. doi: 10.1074/jbc.M801500200. Epub 2008 Jul 23. J Biol Chem. 2008. PMID: 18650424 Free PMC article.
-
Biological functions of miR-29b contribute to positive regulation of osteoblast differentiation.J Biol Chem. 2009 Jun 5;284(23):15676-84. doi: 10.1074/jbc.M809787200. Epub 2009 Apr 2. J Biol Chem. 2009. PMID: 19342382 Free PMC article.
-
High throughput transcriptome profiling of lithium stimulated human mesenchymal stem cells reveals priming towards osteoblastic lineage.PLoS One. 2013;8(1):e55769. doi: 10.1371/journal.pone.0055769. Epub 2013 Jan 30. PLoS One. 2013. PMID: 23383279 Free PMC article.
References
-
- Caunt CJ, Finch AR, Sedgley KR, McArdle CA. Trends Endocrinol. Metab. 2006;17:276–283. - PubMed
-
- Ebisuya M, Kondoh K, Nishida E. J. Cell Sci. 2005;118:2997–3002. - PubMed
-
- Murphy LO, Blenis J. Trends Biochem. Sci. 2006;31:268–275. - PubMed
-
- Murphy LO, Smith S, Chen RH, Fingar DC, Blenis J. Nat. Cell Biol. 2002;4:556–564. - PubMed
-
- Yamamoto T, Ebisuya M, Ashida F, Okamoto K, Yonehara S, Nishida E. Curr. Biol. 2006;16:1171–1182. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

