Theoretical study of sequence-dependent nanopore unzipping of DNA
- PMID: 18178661
- PMCID: PMC2267141
- DOI: 10.1529/biophysj.107.111732
Theoretical study of sequence-dependent nanopore unzipping of DNA
Abstract
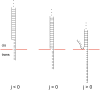
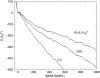
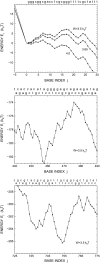
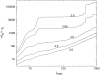
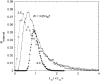
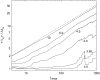
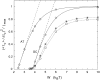
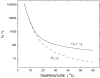
We theoretically investigate the unzipping of DNA electrically driven through a nanometer-size pore. Taking the DNA base sequence explicitly into account, the unpairing and translocation process is described by a biased random walk in a one-dimensional energy landscape determined by the sequential basepair opening. Distributions of translocation times are numerically calculated as a function of applied voltage and temperature. We show that varying these two parameters changes the dynamics from a predominantly diffusive behavior to a dynamics governed by jumps over local energy barriers. The work suggests experimentally studying sequence effects, by comparing the average value and standard deviation of the statistical distribution of translocation times.
Figures

Similar articles
-
Force fluctuations assist nanopore unzipping of DNA.J Phys Condens Matter. 2010 Nov 17;22(45):454122. doi: 10.1088/0953-8984/22/45/454122. Epub 2010 Oct 29. J Phys Condens Matter. 2010. PMID: 21339609
-
Simulations of nanopore formation and phosphatidylserine externalization in lipid membranes subjected to a high-intensity, ultrashort electric pulse.Phys Rev E Stat Nonlin Soft Matter Phys. 2005 Sep;72(3 Pt 1):031902. doi: 10.1103/PhysRevE.72.031902. Epub 2005 Sep 8. Phys Rev E Stat Nonlin Soft Matter Phys. 2005. PMID: 16241477
-
DNA translocation governed by interactions with solid-state nanopores.Biophys J. 2008 Nov 15;95(10):4716-25. doi: 10.1529/biophysj.108.140475. Epub 2008 Aug 15. Biophys J. 2008. PMID: 18708467 Free PMC article.
-
Quantitative analysis of the nanopore translocation dynamics of simple structured polynucleotides.Biophys J. 2012 Jan 4;102(1):85-95. doi: 10.1016/j.bpj.2011.11.4011. Epub 2012 Jan 3. Biophys J. 2012. PMID: 22225801 Free PMC article.
-
Nanopore-based single-molecule DNA analysis.Nanomedicine (Lond). 2007 Aug;2(4):459-81. doi: 10.2217/17435889.2.4.459. Nanomedicine (Lond). 2007. PMID: 17716132 Review.
Cited by
-
Unzipping kinetics of duplex DNA containing oxidized lesions in an α-hemolysin nanopore.J Am Chem Soc. 2012 Jul 4;134(26):11006-11. doi: 10.1021/ja304169n. Epub 2012 Jun 25. J Am Chem Soc. 2012. PMID: 22690806 Free PMC article.
-
Shrinking of Solid-state Nanopores by Direct Thermal Heating.Nanoscale Res Lett. 2011 May 4;6(1):372. doi: 10.1186/1556-276X-6-372. Nanoscale Res Lett. 2011. PMID: 21711885 Free PMC article.
-
Possible scenarios of DNA double-helix unzipping process in single-molecule manipulation experiments.Eur Biophys J. 2018 Dec;47(8):917-924. doi: 10.1007/s00249-018-1313-3. Epub 2018 May 31. Eur Biophys J. 2018. PMID: 29855676
-
Probing DNA base pairing energy profiles using a nanopore.Eur Biophys J. 2009 Feb;38(2):263-9. doi: 10.1007/s00249-008-0372-2. Epub 2008 Oct 3. Eur Biophys J. 2009. PMID: 18836709
-
Overstretching Double-Stranded RNA, Double-Stranded DNA, and RNA-DNA Duplexes.Biophys J. 2019 Aug 6;117(3):509-519. doi: 10.1016/j.bpj.2019.07.003. Epub 2019 Jul 9. Biophys J. 2019. PMID: 31337545 Free PMC article.
References
-
- Sauer-Budge, A. F., J. A. Nyamwanda, D. K. Lubensky, and D. Branton. 2003. Unzipping kinetics of double-stranded DNA in a nanopore. Phys. Rev. Lett. 90:238101. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

