A new method for class prediction based on signed-rank algorithms applied to Affymetrix microarray experiments
- PMID: 18190711
- PMCID: PMC2248160
- DOI: 10.1186/1471-2105-9-16
A new method for class prediction based on signed-rank algorithms applied to Affymetrix microarray experiments
Abstract
Background: The huge amount of data generated by DNA chips is a powerful basis to classify various pathologies. However, constant evolution of microarray technology makes it difficult to mix data from different chip types for class prediction of limited sample populations. Affymetrix(R) technology provides both a quantitative fluorescence signal and a decision (detection call: absent or present) based on signed-rank algorithms applied to several hybridization repeats of each gene, with a per-chip normalization. We developed a new prediction method for class belonging based on the detection call only from recent Affymetrix chip type. Biological data were obtained by hybridization on U133A, U133B and U133Plus 2.0 microarrays of purified normal B cells and cells from three independent groups of multiple myeloma (MM) patients.
Results: After a call-based data reduction step to filter out non class-discriminative probe sets, the gene list obtained was reduced to a predictor with correction for multiple testing by iterative deletion of probe sets that sequentially improve inter-class comparisons and their significance. The error rate of the method was determined using leave-one-out and 5-fold cross-validation. It was successfully applied to (i) determine a sex predictor with the normal donor group classifying gender with no error in all patient groups except for male MM samples with a Y chromosome deletion, (ii) predict the immunoglobulin light and heavy chains expressed by the malignant myeloma clones of the validation group and (iii) predict sex, light and heavy chain nature for every new patient. Finally, this method was shown powerful when compared to the popular classification method Prediction Analysis of Microarray (PAM).
Conclusion: This normalization-free method is routinely used for quality control and correction of collection errors in patient reports to clinicians. It can be easily extended to multiple class prediction suitable with clinical groups, and looks particularly promising through international cooperative projects like the "Microarray Quality Control project of US FDA" MAQC as a predictive classifier for diagnostic, prognostic and response to treatment. Finally, it can be used as a powerful tool to mine published data generated on Affymetrix systems and more generally classify samples with binary feature values.
Figures
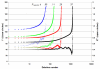
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

