A large scale analysis of protein-protein interactions in the nitrogen-fixing bacterium Mesorhizobium loti
- PMID: 18192278
- PMCID: PMC2650630
- DOI: 10.1093/dnares/dsm028
A large scale analysis of protein-protein interactions in the nitrogen-fixing bacterium Mesorhizobium loti
Abstract
Global viewing of protein-protein interactions (PPIs) is a useful way to assign biological roles to large numbers of proteins predicted by complete genome sequence. Here, we systematically analyzed PPIs in the nitrogen-fixing soil bacterium Mesorhizobium loti using a modified high-throughput yeast two-hybrid system. The aims of this study are primarily on the providing functional clues to M. loti proteins that are relevant to symbiotic nitrogen fixation and conserved in other rhizobium species, especially proteins with regulatory functions and unannotated proteins. By the screening of 1542 genes as bait, 3121 independent interactions involving 1804 proteins (24% of the total protein coding genes) were identified and each interaction was evaluated using an interaction generality (IG) measure and the general features of the interacting partners. Most PPIs detected in this study are novel interactions revealing potential functional relationships between genes for symbiotic nitrogen fixation and signal transduction. Furthermore, we have predicted the putative functions of unannotated proteins through their interactions with known proteins. The results described here represent new insight into protein network of M. loti and provide useful experimental clues to elucidate the biological function of rhizobial genes that can not be assigned directly from their genomic sequence.
Figures
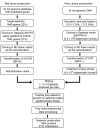

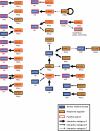
References
-
- Kaneko T., Nakamura Y., Sato S., et al. Complete genome structure of the nitrogen-fixing symbiotic bacterium Mesorhizobium loti. DNA Res. 2000;31:331–338. - PubMed
-
- Kaneko T., Nakamura Y., Sato S., et al. Complete genomic sequence of nitrogen-fixing symbiotic bacterium Bradyrhizobium japonicum USDA110. DNA Res. 2002;9:189–197. - PubMed
-
- Galibert F., Finan T. M., Long S. R., et al. The composite genome of the legume symbiont Sinorhizobium meliloti. Science. 2001;293:668–672. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

