Identification of Campylobacter jejuni genes involved in the response to acidic pH and stomach transit
- PMID: 18192414
- PMCID: PMC2258634
- DOI: 10.1128/AEM.01507-07
Identification of Campylobacter jejuni genes involved in the response to acidic pH and stomach transit
Abstract
Campylobacter jejuni causes food- and waterborne gastroenteritis, and as such it must survive passage through the stomach in order to reach the gastrointestinal tract. While little is known about how C. jejuni survives transit through the stomach, its low infectious dose suggests it is well equipped to sense and respond to acid shock. In this study, the transcriptional profile of C. jejuni NCTC 11168 was obtained after the organism was exposed to in vitro and in vivo (piglet stomach) acid shock. The observed down-regulation of genes encoding ribosomal proteins likely reflects the need to reshuffle energy toward the expression of components required for survival. Acid shock also caused C. jejuni to up-regulate genes involved in stress responses. These included heat shock genes as well as genes involved in the response to oxidative and nitrosative stress. A role for the chaperone clpB in acid resistance was confirmed in vitro. Some genes showed expression patterns that were markedly different in vivo and in vitro, which likely reflects the complexity of the in vivo environment. For instance, transit through the stomach was characterized by up-regulation of genes that encode products that are involved in the use of nitrite as a terminal electron acceptor and down-regulation of genes that are involved in capsular polysaccharide expression. In conclusion, this study has enabled us to understand how C. jejuni modulates gene expression in response to acid shock in vitro and to correlate this with gene expression profiles of C. jejuni as it transits through the host stomach.
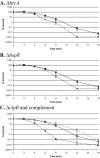
Figures





Similar articles
-
Identification of Campylobacter jejuni genes contributing to acid adaptation by transcriptional profiling and genome-wide mutagenesis.Appl Environ Microbiol. 2008 Mar;74(5):1598-612. doi: 10.1128/AEM.01508-07. Epub 2008 Jan 11. Appl Environ Microbiol. 2008. PMID: 18192408 Free PMC article.
-
Gene expression profile of Campylobacter jejuni in response to growth temperature variation.J Bacteriol. 2003 Mar;185(6):2009-16. doi: 10.1128/JB.185.6.2009-2016.2003. J Bacteriol. 2003. PMID: 12618466 Free PMC article.
-
Diverse roles for HspR in Campylobacter jejuni revealed by the proteome, transcriptome and phenotypic characterization of an hspR mutant.Microbiology (Reading). 2005 Mar;151(Pt 3):905-915. doi: 10.1099/mic.0.27513-0. Microbiology (Reading). 2005. PMID: 15758235
-
CmeR functions as a pleiotropic regulator and is required for optimal colonization of Campylobacter jejuni in vivo.J Bacteriol. 2008 Mar;190(6):1879-90. doi: 10.1128/JB.01796-07. Epub 2008 Jan 4. J Bacteriol. 2008. PMID: 18178742 Free PMC article.
-
The Campylobacter jejuni Ferric Uptake Regulator Promotes Acid Survival and Cross-Protection against Oxidative Stress.Infect Immun. 2016 Apr 22;84(5):1287-1300. doi: 10.1128/IAI.01377-15. Print 2016 May. Infect Immun. 2016. PMID: 26883589 Free PMC article.
Cited by
-
Cj1386, an atypical hemin-binding protein, mediates hemin trafficking to KatA in Campylobacter jejuni.J Bacteriol. 2015 Mar;197(5):1002-11. doi: 10.1128/JB.02346-14. Epub 2014 Dec 29. J Bacteriol. 2015. PMID: 25548249 Free PMC article.
-
Polyphosphate kinase 2: a novel determinant of stress responses and pathogenesis in Campylobacter jejuni.PLoS One. 2010 Aug 17;5(8):e12142. doi: 10.1371/journal.pone.0012142. PLoS One. 2010. PMID: 20808906 Free PMC article.
-
Comparative genomics analysis and virulence-related factors in novel Aliarcobacter faecis and Aliarcobacter lanthieri species identified as potential opportunistic pathogens.BMC Genomics. 2022 Jun 27;23(1):471. doi: 10.1186/s12864-022-08663-w. BMC Genomics. 2022. PMID: 35761183 Free PMC article.
-
Advances in Campylobacter biology and implications for biotechnological applications.Microb Biotechnol. 2010 May;3(3):242-58. doi: 10.1111/j.1751-7915.2009.00118.x. Epub 2009 May 19. Microb Biotechnol. 2010. PMID: 21255325 Free PMC article. Review.
-
Functional insights into the interplay between DNA interaction and metal coordination in ferric uptake regulators.Sci Rep. 2018 May 8;8(1):7140. doi: 10.1038/s41598-018-25157-6. Sci Rep. 2018. PMID: 29739988 Free PMC article.
References
-
- Allan, E., C. L. Clayton, A. McLaren, D. M. Wallace, and B. W. Wren. 2001. Characterization of the low-pH responses of Helicobacter pylori using genomic DNA arrays. Microbiology 147:2285-2292. - PubMed
-
- Andersen, M. T., L. Brondsted, B. M. Pearson, F. Mulholland, M. Parker, C. Pin, J. M. Wells, and H. Ingmer. 2005. Diverse roles for HspR in Campylobacter jejuni revealed by the proteome, transcriptome and phenotypic characterization of an hspR mutant. Microbiology 151:905-915. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

