Identification of an OsPR10a promoter region responsive to salicylic acid
- PMID: 18193274
- PMCID: PMC2270913
- DOI: 10.1007/s00425-007-0687-8
Identification of an OsPR10a promoter region responsive to salicylic acid
Abstract
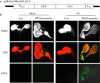
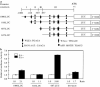
Orysa sativa pathogenesis-related protein 10a (OsPR10a) was induced by pathogens, salicylic acid (SA), jasmonic acid (JA), ethephon, abscisic acid (ABA), and NaCl. We tried to analyze the OsPR10a promoter to investigate the transcriptional regulation of OsPR10a by SA. We demonstrated the inducibility of OsPR10a promoter by SA using transgenic Arabidopsis carrying OsPR10a:GFP as well as by transient expression assays in rice. To further identify the promoter region responsible for its induction by SA, four different deletions of the OsPR10a promoter were made, and their activities were measured by transient assays. The construct containing 687-bp OsPR10a promoter from its start codon exhibited a six-fold increase of induction compared to the control in response to SA. Mutation in the W-box like element 1 (WLE 1) between 687 and 637-bp from TGACA to TGAAA completely abolished induction of the OsPR10a promoter by SA, indicating that the WLE 1 between -687 and -637 of OsPR10a promoter is important in SA-mediated OsPR10a expression. We show for the first time that the W-box like element plays a role in SA mediated PR gene expression.
Figures




Similar articles
-
Molecular characterization of Oryza sativa WRKY6, which binds to W-box-like element 1 of the Oryza sativa pathogenesis-related (PR) 10a promoter and confers reduced susceptibility to pathogens.New Phytol. 2015 Nov;208(3):846-59. doi: 10.1111/nph.13516. Epub 2015 Jun 17. New Phytol. 2015. PMID: 26083148
-
A W-box is required for full expression of the SA-responsive gene SFR2.Gene. 2005 Jan 3;344:181-92. doi: 10.1016/j.gene.2004.09.016. Epub 2004 Nov 19. Gene. 2005. PMID: 15656984
-
Roles of salicylic acid-responsive cis-acting elements and W-boxes in salicylic acid induction of VCH3 promoter in transgenic tobaccos.Acta Biochim Biophys Sin (Shanghai). 2006 Jan;38(1):46-52. doi: 10.1111/j.1745-7270.2006.00126.x. Acta Biochim Biophys Sin (Shanghai). 2006. PMID: 16395526
-
Salicylic acid and a chitin elicitor both control expression of the CAD1 gene involved in the plant immunity of Arabidopsis.Biosci Biotechnol Biochem. 2006 Sep;70(9):2042-8. doi: 10.1271/bbb.50700. Epub 2006 Sep 7. Biosci Biotechnol Biochem. 2006. PMID: 16960394
-
Functional analysis of a TGA factor-binding site located in the promoter region controlling salicylic acid-induced NIMIN-1 expression in Arabidopsis.Genet Mol Res. 2010 Feb 2;9(1):167-75. doi: 10.4238/vol9-1gmr704. Genet Mol Res. 2010. PMID: 20198573
Cited by
-
Overexpression of a constitutively active truncated form of OsCDPK1 confers disease resistance by affecting OsPR10a expression in rice.Sci Rep. 2018 Jan 10;8(1):403. doi: 10.1038/s41598-017-18829-2. Sci Rep. 2018. PMID: 29321675 Free PMC article.
-
Association of genetic variants in the Sirt1 and Nrf2 genes with the risk of metabolic syndrome in a Chinese Han population.BMC Endocr Disord. 2022 Apr 1;22(1):84. doi: 10.1186/s12902-022-00965-0. BMC Endocr Disord. 2022. PMID: 35365152 Free PMC article.
-
Inducible Enrichment of Osa-miR1432 Confers Rice Bacterial Blight Resistance through Suppressing OsCaML2.Int J Mol Sci. 2021 Oct 21;22(21):11367. doi: 10.3390/ijms222111367. Int J Mol Sci. 2021. PMID: 34768797 Free PMC article.
-
Cytochrome P450 monooxygenases as reporters for circadian-regulated pathways.Plant Physiol. 2009 Jun;150(2):858-78. doi: 10.1104/pp.108.130757. Epub 2009 Apr 22. Plant Physiol. 2009. PMID: 19386812 Free PMC article.
-
The OsICS1 is directly regulated by OsWRKY6 and increases resistance against Xanthomonas oryzae pv. oryzae.Planta. 2024 Apr 17;259(6):124. doi: 10.1007/s00425-024-04405-2. Planta. 2024. PMID: 38630137
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

