Identification of the minus-dominance gene ortholog in the mating-type locus of Gonium pectorale
- PMID: 18202374
- PMCID: PMC2206078
- DOI: 10.1534/genetics.107.078618
Identification of the minus-dominance gene ortholog in the mating-type locus of Gonium pectorale
Abstract
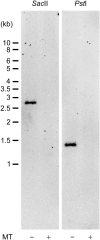
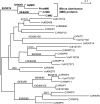
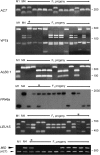
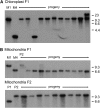
The evolution of anisogamy/oogamy in the colonial Volvocales might have occurred in an ancestral isogamous colonial organism like Gonium pectorale. The unicellular, close relative Chlamydomonas reinhardtii has a mating-type (MT) locus harboring several mating-type-specific genes, including one involved in mating-type determination and another involved in the function of the tubular mating structure in only one of the two isogametes. In this study, as the first step in identifying the G. pectorale MT locus, we isolated from G. pectorale the ortholog of the C. reinhardtii mating-type-determining minus-dominance (CrMID) gene, which is localized only in the MT- locus. 3'- and 5'-RACE RT-PCR using degenerate primers identified a CrMID-orthologous 164-amino-acid coding gene (GpMID) containing a leucine-zipper RWP-RK domain near the C-terminal, as is the case with CrMID. Genomic Southern blot analysis showed that GpMID was coded only in the minus strain of G. pectorale. RT-PCR revealed that GpMID expression increased during nitrogen starvation. Analysis of F1 progeny suggested that GpMID and isopropylmalate dehydratase LEU1S are tightly linked, suggesting that they are harbored in a chromosomal region under recombinational suppression that is comparable to the C. reinhardtii MT locus. However, two other genes present in the C. reinhardtii MT locus are not linked to the G. pectorale LEU1S/MID, suggesting that the gene content of the volvocalean MT loci is not static over time. Inheritance of chloroplast and mitochondria genomes in G. pectorale is uniparental from the plus and minus parents, respectively, as is also the case in C. reinhardtii.
Figures




References
-
- Adams, C. R., K. A. Stamer, J. K. Miller, J. G. McNally, M. M. Kirk et al., 1990. Patterns of organellar and nuclear inheritance among progeny of two geographically isolated strains of Volvox carteri. Curr. Genet. 18: 141–153. - PubMed
-
- Burge, C., and S. Karlin, 1997. Prediction of complete gene structures in human genomic DNA. J. Mol. Biol. 268: 78–94. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Research Materials

