WRN controls formation of extrachromosomal telomeric circles and is required for TRF2DeltaB-mediated telomere shortening
- PMID: 18212065
- PMCID: PMC2268394
- DOI: 10.1128/MCB.01364-07
WRN controls formation of extrachromosomal telomeric circles and is required for TRF2DeltaB-mediated telomere shortening
Abstract
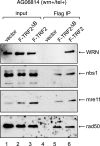
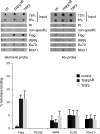
Telomere dysfunction has been proposed to contribute to the pathogenesis of Werner syndrome (WS), a premature-aging disorder. The WS protein WRN binds TRF2, a telomere-specific factor that protects chromosome ends. TRF2 possesses an amino-terminal domain that plays an essential role in preventing telomere shortening, as expression of TRF2(DeltaB), which lacks this domain, leads to the formation of telomeric circles, telomere shortening, and cell senescence. Our data show that the TRF2(DeltaB)-induced telomeric-loop homologous-recombination pathway requires WRN helicase. In addition, we show that WRN represses the formation of spontaneous telomeric circles, as demonstrated by the increased levels of telomeric circles observed in telomerase-positive WS fibroblasts. The mechanism of circle formation in WS cells does not involve XRCC3 function. Circle formation in WS cells is reduced by reconstitution with wild-type WRN but not mutant forms lacking either exonuclease or helicase activity, demonstrating that both enzymatic activities of WRN are required to suppress telomeric-circle formation in normal cells expressing telomerase reverse transcriptase. Thus, WRN has a key protective function at telomeres which influences telomere topology and inhibits accelerated attrition of telomeres.
Figures


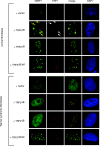
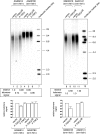
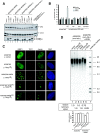
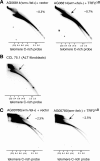

Similar articles
-
TRF2 recruits the Werner syndrome (WRN) exonuclease for processing of telomeric DNA.Oncogene. 2004 Jan 8;23(1):149-56. doi: 10.1038/sj.onc.1206906. Oncogene. 2004. PMID: 14712220
-
Sequence-specific processing of telomeric 3' overhangs by the Werner syndrome protein exonuclease activity.Aging (Albany NY). 2009 Mar 17;1(3):289-302. doi: 10.18632/aging.100032. Aging (Albany NY). 2009. PMID: 20157518 Free PMC article.
-
The Werner syndrome helicase and exonuclease cooperate to resolve telomeric D loops in a manner regulated by TRF1 and TRF2.Mol Cell. 2004 Jun 18;14(6):763-74. doi: 10.1016/j.molcel.2004.05.023. Mol Cell. 2004. PMID: 15200954
-
Telomere ResQue and preservation--roles for the Werner syndrome protein and other RecQ helicases.Mech Ageing Dev. 2008 Jan-Feb;129(1-2):79-90. doi: 10.1016/j.mad.2007.10.007. Epub 2007 Oct 30. Mech Ageing Dev. 2008. PMID: 18054793 Review.
-
Role of Werner syndrome gene product helicase in carcinogenesis and in resistance to genotoxins by cancer cells.Cancer Sci. 2008 May;99(5):843-8. doi: 10.1111/j.1349-7006.2008.00778.x. Epub 2008 Feb 26. Cancer Sci. 2008. PMID: 18312465 Free PMC article. Review.
Cited by
-
Telomeric protein TRF2 protects Holliday junctions with telomeric arms from displacement by the Werner syndrome helicase.Nucleic Acids Res. 2010 Jul;38(12):3984-98. doi: 10.1093/nar/gkq144. Epub 2010 Mar 9. Nucleic Acids Res. 2010. PMID: 20215438 Free PMC article.
-
Ku circles the telomere?Aging (Albany NY). 2011 Apr;3(4):344-5. doi: 10.18632/aging.100321. Aging (Albany NY). 2011. PMID: 21732567 Free PMC article. No abstract available.
-
Replication protein A stimulates the Werner syndrome protein branch migration activity.J Biol Chem. 2009 Dec 11;284(50):34682-91. doi: 10.1074/jbc.M109.049031. Epub 2009 Oct 7. J Biol Chem. 2009. PMID: 19812417 Free PMC article.
-
Ku86 represses lethal telomere deletion events in human somatic cells.Proc Natl Acad Sci U S A. 2009 Jul 28;106(30):12430-5. doi: 10.1073/pnas.0903362106. Epub 2009 Jul 6. Proc Natl Acad Sci U S A. 2009. PMID: 19581589 Free PMC article.
-
Acrolein-exposed normal human lung fibroblasts in vitro: cellular senescence, enhanced telomere erosion, and degradation of Werner's syndrome protein.Environ Health Perspect. 2014 Sep;122(9):955-62. doi: 10.1289/ehp.1306911. Epub 2014 Apr 18. Environ Health Perspect. 2014. PMID: 24747221 Free PMC article.
References
-
- Bilaud, T., C. Brun, K. Ancelin, C. E. Koering, T. Laroche, and E. Gilson. 1997. Telomeric localization of TRF2, a novel human telobox protein. Nat. Genet. 17236-239. - PubMed
-
- Brewer, B. J., and W. L. Fangman. 1987. The localization of replication origins on ARS plasmids in S. cerevisiae. Cell 51463-471. - PubMed
-
- Broccoli, D., A. Smogorzewska, L. Chong, and T. de Lange. 1997. Human telomeres contain two distinct Myb-related proteins, TRF1 and TRF2. Nat. Genet. 17231-235. - PubMed
-
- Chong, L., B. van Steensel, D. Broccoli, H. Erdjument-Bromage, J. Hanish, P. Tempst, and T. de Lange. 1995. A human telomeric protein. Science 2701663-1667. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous
