Stress-dependent dynamics of global chromatin remodeling in yeast: dual role for SWI/SNF in the heat shock stress response
- PMID: 18212068
- PMCID: PMC2268435
- DOI: 10.1128/MCB.01659-07
Stress-dependent dynamics of global chromatin remodeling in yeast: dual role for SWI/SNF in the heat shock stress response
Abstract
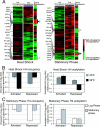
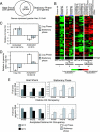
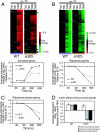
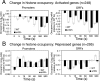
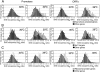
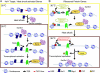
Although chromatin structure is known to affect transcriptional activity, it is not clear how broadly patterns of changes in histone modifications and nucleosome occupancy affect the dynamic regulation of transcription in response to perturbations. The identity and role of chromatin remodelers that mediate some of these changes are also unclear. Here, we performed temporal genome-wide analyses of gene expression, nucleosome occupancy, and histone H4 acetylation during the response of yeast (Saccharomyces cerevisiae) to different stresses and report several findings. First, a large class of predominantly ribosomal protein genes, whose transcription was repressed during both heat shock and stationary phase, showed strikingly contrasting histone acetylation patterns. Second, the SWI/SNF complex was required for normal activation as well as repression of genes during heat shock, and loss of SWI/SNF delayed chromatin remodeling at the promoters of activated genes. Third, Snf2 was recruited to ribosomal protein genes and Hsf1 target genes, and its occupancy of this large set of genes was altered during heat shock. Our results suggest a broad and direct dual role for SWI/SNF in chromatin remodeling, during heat shock activation as well as repression, at promoters and coding regions.
Figures
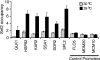
Similar articles
-
Mediator, TATA-binding protein, and RNA polymerase II contribute to low histone occupancy at active gene promoters in yeast.J Biol Chem. 2014 May 23;289(21):14981-95. doi: 10.1074/jbc.M113.529354. Epub 2014 Apr 11. J Biol Chem. 2014. PMID: 24727477 Free PMC article.
-
Genetic and epigenetic determinants establish a continuum of Hsf1 occupancy and activity across the yeast genome.Mol Biol Cell. 2018 Dec 15;29(26):3168-3182. doi: 10.1091/mbc.E18-06-0353. Epub 2018 Oct 17. Mol Biol Cell. 2018. PMID: 30332327 Free PMC article.
-
Different requirements of the SWI/SNF complex for robust nucleosome displacement at promoters of heat shock factor and Msn2- and Msn4-regulated heat shock genes.Mol Cell Biol. 2008 Feb;28(4):1207-17. doi: 10.1128/MCB.01069-07. Epub 2007 Dec 10. Mol Cell Biol. 2008. PMID: 18070923 Free PMC article.
-
Yeast chromatin remodeling complexes and their roles in transcription.Curr Genet. 2020 Aug;66(4):657-670. doi: 10.1007/s00294-020-01072-0. Epub 2020 Apr 1. Curr Genet. 2020. PMID: 32239283 Review.
-
The SWI/SNF chromatin remodeling complex: a critical regulator of metabolism.Biochem Soc Trans. 2024 Jun 26;52(3):1327-1337. doi: 10.1042/BST20231141. Biochem Soc Trans. 2024. PMID: 38666605 Free PMC article. Review.
Cited by
-
Distinct functions of three chromatin remodelers in activator binding and preinitiation complex assembly.PLoS Genet. 2022 Jul 6;18(7):e1010277. doi: 10.1371/journal.pgen.1010277. eCollection 2022 Jul. PLoS Genet. 2022. PMID: 35793348 Free PMC article.
-
Regulation of alternative polyadenylation in the yeast Saccharomyces cerevisiae by histone H3K4 and H3K36 methyltransferases.Nucleic Acids Res. 2020 Jun 4;48(10):5407-5425. doi: 10.1093/nar/gkaa292. Nucleic Acids Res. 2020. PMID: 32356874 Free PMC article.
-
Genome information processing by the INO80 chromatin remodeler positions nucleosomes.Nat Commun. 2021 May 28;12(1):3231. doi: 10.1038/s41467-021-23016-z. Nat Commun. 2021. PMID: 34050142 Free PMC article.
-
Control of Gene Expression via the Yeast CWI Pathway.Int J Mol Sci. 2022 Feb 4;23(3):1791. doi: 10.3390/ijms23031791. Int J Mol Sci. 2022. PMID: 35163713 Free PMC article. Review.
-
Swi/Snf chromatin remodeling regulates transcriptional interference and gene repression.Mol Cell. 2024 Aug 22;84(16):3080-3097.e9. doi: 10.1016/j.molcel.2024.06.029. Epub 2024 Jul 22. Mol Cell. 2024. PMID: 39043178 Free PMC article.
References
-
- Boy-Marcotte, E., G. Lagniel, M. Perrot, F. Bussereau, A. Boudsocq, M. Jacquet, and J. Labarre. 1999. The heat shock response in yeast: differential regulations and contributions of the Msn2p/Msn4p and Hsf1p regulons. Mol. Microbiol. 33274-283. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
