Dynamics of the yeast transcriptome during wine fermentation reveals a novel fermentation stress response
- PMID: 18215224
- PMCID: PMC5065349
- DOI: 10.1111/j.1567-1364.2007.00338.x
Dynamics of the yeast transcriptome during wine fermentation reveals a novel fermentation stress response
Abstract
In this study, genome-wide expression analyses were used to study the response of Saccharomyces cerevisiae to stress throughout a 15-day wine fermentation. Forty per cent of the yeast genome significantly changed expression levels to mediate long-term adaptation to fermenting grape must. Among the genes that changed expression levels, a group of 223 genes was identified, which was designated as fermentation stress response (FSR) genes that were dramatically induced at various points during fermentation. FSR genes sustain high levels of induction up to the final time point and exhibited changes in expression levels ranging from four- to 80-fold. The FSR is novel; 62% of the genes involved have not been implicated in global stress responses and 28% of the FSR genes have no functional annotation. Genes involved in respiratory metabolism and gluconeogenesis were expressed during fermentation despite the presence of high concentrations of glucose. Ethanol, rather than nutrient depletion, seems to be responsible for entry of yeast cells into the stationary phase.
Figures


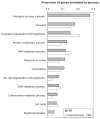


References
-
- Ahuatzi D, Riera A, Pelaez R, Herrero P, Moreno F. Hxk2 regulates the phosphorylation state of Mig1 and therefore its nucleocytoplasmic distribution. J Biol Chem. 2007;282:4485–4493. - PubMed
-
- Alexandre H, Ansanay-Galeote V, Dequin S, Blondin B. Global gene expression during short-term ethanol stress in Saccharomyces cerevisiae. FEBS Lett. 2001;498:98–103. - PubMed
-
- Benjamini Y, Hochberg Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Statist Soc B. 1995;57:289–300.
-
- Brewster JL, de Valoir T, Dwyer ND, Winter E, Gustin MC. An osmosensing signal transduction pathway in yeast. Science. 1993;259:1760–1763. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

