Dynactin function in mitotic spindle positioning
- PMID: 18221362
- PMCID: PMC2367371
- DOI: 10.1111/j.1600-0854.2008.00710.x
Dynactin function in mitotic spindle positioning
Abstract
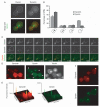
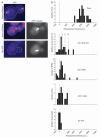
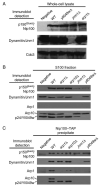
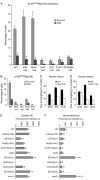
Dynactin is a multisubunit protein complex necessary for dynein function. Here, we investigated the function of dynactin in budding yeast. Loss of dynactin impaired movement and positioning of the mitotic spindle, similar to loss of dynein. Dynactin subunits required for function included p150(Glued), dynamitin, actin-related protein (Arp) 1 and p24. Arp10 and capping protein were dispensable, even in combination. All dynactin subunits tested localized to dynamic plus ends of cytoplasmic microtubules, to stationary foci on the cell cortex and to spindle pole bodies. The number of molecules of dynactin in those locations was small, less than five. In the absence of dynactin, dynein accumulated at plus ends and did not appear at the cell cortex, consistent with a role for dynactin in offloading dynein from the plus end to the cortex. Dynein at the plus end was necessary for dynactin plus-end targeting. p150(Glued) was the only dynactin subunit sufficient for plus-end targeting. Interactions among the subunits support a molecular model that resembles the current model for brain dynactin in many respects; however, three subunits at the pointed end of brain dynactin appear to be absent from yeast.
Figures


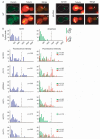

Similar articles
-
The role of the lissencephaly protein Pac1 during nuclear migration in budding yeast.J Cell Biol. 2003 Feb 3;160(3):355-64. doi: 10.1083/jcb.200209022. J Cell Biol. 2003. PMID: 12566428 Free PMC article.
-
Dynein-driven mitotic spindle positioning restricted to anaphase by She1p inhibition of dynactin recruitment.Mol Biol Cell. 2009 Jul;20(13):3003-11. doi: 10.1091/mbc.e09-03-0186. Epub 2009 Apr 29. Mol Biol Cell. 2009. PMID: 19403691 Free PMC article.
-
Dynein is regulated by the stability of its microtubule track.J Cell Biol. 2017 Jul 3;216(7):2047-2058. doi: 10.1083/jcb.201611105. Epub 2017 Jun 1. J Cell Biol. 2017. PMID: 28572117 Free PMC article.
-
The role of the dynactin complex in intracellular motility.Int Rev Cytol. 1998;182:69-109. doi: 10.1016/s0074-7696(08)62168-3. Int Rev Cytol. 1998. PMID: 9522459 Review.
-
[Dynein and dynactin as organizers of the system of cell microtubules].Ontogenez. 2006 Sep-Oct;37(5):323-39. Ontogenez. 2006. PMID: 17066975 Review. Russian.
Cited by
-
Conserved roles for the dynein intermediate chain and Ndel1 in assembly and activation of dynein.Nat Commun. 2023 Sep 20;14(1):5833. doi: 10.1038/s41467-023-41466-5. Nat Commun. 2023. PMID: 37730751 Free PMC article.
-
An RNAi screen for genes that affect nuclear morphology in Caenorhabditis elegans reveals the involvement of unexpected processes.G3 (Bethesda). 2021 Oct 19;11(11):jkab264. doi: 10.1093/g3journal/jkab264. G3 (Bethesda). 2021. PMID: 34849797 Free PMC article.
-
Pac1/LIS1 stabilizes an uninhibited conformation of dynein to coordinate its localization and activity.Nat Cell Biol. 2020 May;22(5):559-569. doi: 10.1038/s41556-020-0492-1. Epub 2020 Apr 27. Nat Cell Biol. 2020. PMID: 32341548 Free PMC article.
-
VezA/vezatin facilitates proper assembly of the dynactin complex in vivo.bioRxiv [Preprint]. 2024 Apr 20:2024.04.19.590248. doi: 10.1101/2024.04.19.590248. bioRxiv. 2024. Update in: Cell Rep. 2024 Nov 26;43(11):114943. doi: 10.1016/j.celrep.2024.114943. PMID: 38659795 Free PMC article. Updated. Preprint.
-
A novel mechanism of bulk cytoplasmic transport by cortical dynein in Drosophila ovary.Elife. 2022 Feb 16;11:e75538. doi: 10.7554/eLife.75538. Elife. 2022. PMID: 35170428 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

