Degradation of 4-fluorophenol by Arthrobacter sp. strain IF1
- PMID: 18228015
- PMCID: PMC2266783
- DOI: 10.1007/s00253-008-1343-3
Degradation of 4-fluorophenol by Arthrobacter sp. strain IF1
Abstract
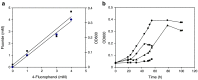
A Gram-positive bacterial strain capable of aerobic biodegradation of 4-fluorophenol (4-FP) as the sole source of carbon and energy was isolated by selective enrichment from soil samples collected near an industrial site. The organism, designated strain IF1, was identified as a member of the genus Arthrobacter on the basis of 16S ribosomal RNA gene sequence analysis. Arthrobacter strain IF1 was able to mineralize 4-FP up to concentrations of 5 mM in batch culture. Stoichiometric release of fluoride ions was observed, suggesting that there is no formation of halogenated dead-end products during 4-FP metabolism. The degradative pathway of 4-FP was investigated using enzyme assays and identification of intermediates by gas chromatography (GC), GC-mass spectrometry (MS), high-performance liquid chromatography, and liquid chromatography-MS. Cell-free extracts of 4-FP-grown cells contained no activity for catechol 1,2-dioxygenase or catechol 2,3-dioxygenase, which indicates that the pathway does not proceed through a catechol intermediate. Cells grown on 4-FP oxidized 4-FP, hydroquinone, and hydroxyquinol but not 4-fluorocatechol. During 4-FP metabolism, hydroquinone accumulated as a product. Hydroquinone could be converted to hydroxyquinol, which was further transformed into maleylacetic acid and beta-ketoadipic acid. These results indicate that the biodegradation of 4-FP starts with a 4-FP monooxygenase reaction that yields benzoquinone, which is reduced to hydroquinone and further metabolized via the beta-ketoadipic acid pathway.
Figures



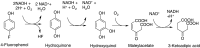
References
-
- Boersma MG, Dinarieva TY, Middelhoven WJ, van Berkel WJ, Doran J, Vervoort J, Rietjens IM. 19F nuclear magnetic resonance as a tool to investigate microbial degradation of fluorophenols to fluorocatechols and fluoromuconates. Appl Environ Microbiol. 1998;64:1256–1263. doi: 10.1128/AEM.64.4.1256-1263.1998. - DOI - PMC - PubMed
-
- Bondar VS, Boersma MG, van Berkel WJ, Finkelstein ZI, Golovlev EL, Baskunov BP, Vervoort J, Golovleva LA, Rietjens IM. Preferential oxidative dehalogenation upon conversion of 2-halophenols by Rhodococcus opacus 1G. FEMS Microbiol Lett. 1999;181:73–82. doi: 10.1111/j.1574-6968.1999.tb08828.x. - DOI - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

