Robust SNP genotyping by multiplex PCR and arrayed primer extension
- PMID: 18237385
- PMCID: PMC2266772
- DOI: 10.1186/1755-8794-1-5
Robust SNP genotyping by multiplex PCR and arrayed primer extension
Abstract
Background: Arrayed primer extension (APEX) is a microarray-based rapid minisequencing methodology that may have utility in 'personalized medicine' applications that involve genetic diagnostics of single nucleotide polymorphisms (SNPs). However, to date there have been few reports that objectively evaluate the assay completion rate, call rate and accuracy of APEX. We have further developed robust assay design, chemistry and analysis methodologies, and have sought to determine how effective APEX is in comparison to leading 'gold-standard' genotyping platforms. Our methods have been tested against industry-leading technologies in two blinded experiments based on Coriell DNA samples and SNP genotype data from the International HapMap Project.
Results: In the first experiment, we genotyped 50 SNPs across the entire 270 HapMap Coriell DNA sample set. For each Coriell sample, DNA template was amplified in a total of 7 multiplex PCRs prior to genotyping. We obtained good results for 41 of the SNPs, with 99.8% genotype concordance with HapMap data, at an automated call rate of 94.9% (not including the 9 failed SNPs). In the second experiment, involving modifications to the initial DNA amplification so that a single 50-plex PCR could be achieved, genotyping of the same 50 SNPs across each of 49 randomly chosen Coriell DNA samples allowed extremely robust 50-plex genotyping from as little as 5 ng of DNA, with 100% assay completion rate, 100% call rate and >99.9% accuracy.
Conclusion: We have shown our methods to be effective for robust multiplex SNP genotyping using APEX, with 100% call rate and >99.9% accuracy. We believe that such methodology may be useful in future point-of-care clinical diagnostic applications where accuracy and call rate are both paramount.
Figures
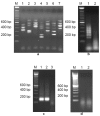


Similar articles
-
Dynamic variable selection in SNP genotype autocalling from APEX microarray data.BMC Bioinformatics. 2006 Nov 30;7:521. doi: 10.1186/1471-2105-7-521. BMC Bioinformatics. 2006. PMID: 17137502 Free PMC article.
-
A novel amplification strategy for genotyping with liquid chromatography-electrospray ionization mass spectrometry.Analyst. 2012 Nov 21;137(22):5325-33. doi: 10.1039/c2an35440c. Analyst. 2012. PMID: 23034565
-
Multiplex automated primer extension analysis: simultaneous genotyping of several polymorphisms.Biotechniques. 2001 Dec;31(6):1374-80. doi: 10.2144/01316md05. Biotechniques. 2001. PMID: 11768667
-
[Approaches for SNP genotyping].Yi Chuan. 2006 Jan;28(1):117-26. Yi Chuan. 2006. PMID: 16469727 Review. Chinese.
-
Enabling large-scale pharmacogenetic studies by high-throughput mutation detection and genotyping technologies.Clin Chem. 2001 Feb;47(2):164-72. Clin Chem. 2001. PMID: 11159763 Review.
Cited by
-
Large-scale development of cost-effective SNP marker assays for diversity assessment and genetic mapping in chickpea and comparative mapping in legumes.Plant Biotechnol J. 2012 Aug;10(6):716-32. doi: 10.1111/j.1467-7652.2012.00710.x. Epub 2012 Jun 16. Plant Biotechnol J. 2012. PMID: 22703242 Free PMC article.
-
Clinical application of high throughput molecular screening techniques for pharmacogenomics.Pharmgenomics Pers Med. 2011;4:109-21. doi: 10.2147/PGPM.S15302. Epub 2011 Sep 8. Pharmgenomics Pers Med. 2011. PMID: 23226057 Free PMC article.
-
Advances in ligase chain reaction and ligation-based amplifications for genotyping assays: Detection and applications.Mutat Res Rev Mutat Res. 2017 Jul;773:66-90. doi: 10.1016/j.mrrev.2017.05.001. Epub 2017 May 2. Mutat Res Rev Mutat Res. 2017. PMID: 28927538 Free PMC article. Review.
References
-
- Hirschhorn JN, Sklar P, Lindblad-Toh K, Lim YM, Ruiz-Gutierrez M, Bolk S, Langhorst B, Schaffner S, Winchester E, Lander ES. SBE-TAGS: an array-based method for efficient single-nucleotide polymorphism genotyping. Proc Natl Acad Sci U S A. 2000;97:12164–12169. doi: 10.1073/pnas.210394597. - DOI - PMC - PubMed
-
- Oliphant A, Barker DL, Stuelpnagel JR, Chee MS. BeadArray technology: enabling an accurate, cost-effective approach to high-throughput genotyping. Biotechniques. 2002;Suppl:56–8, 60-1. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

