Mutations in the Drosophila mitochondrial tRNA amidotransferase, bene/gatA, cause growth defects in mitotic and endoreplicating tissues
- PMID: 18245325
- PMCID: PMC2248357
- DOI: 10.1534/genetics.107.084376
Mutations in the Drosophila mitochondrial tRNA amidotransferase, bene/gatA, cause growth defects in mitotic and endoreplicating tissues
Abstract
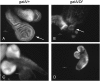
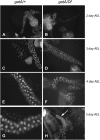
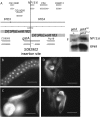
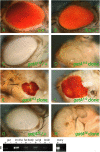
Rapid larval growth is essential in the development of most metazoans. In this article, we show that bene, a gene previously identified on the basis of its oogenesis defects, is also required for larval growth and viability. We show that all bene alleles disrupt gatA, which encodes the Drosophila homolog of glutamyl-tRNA(Gln) amidotransferase subunit A (GatA). bene alleles are now referred to as gatA. GatA proteins are highly conserved throughout eukaryotes and many prokaryotes. These enzymes are required for proper translation of the proteins encoded by the mitochondrial genome and by many eubacterial genomes. Mitotic and endoreplicating tissues in Drosophila gatA loss-of-function mutants grow slowly and never achieve wild-type size, and gatA larvae die before pupariation. gatA mutant eye clones exhibit growth and differentiation defects, indicating that gatA expression is required cell autonomously for normal growth. The gatA gene is widely expressed in mitotic and endoreplicating tissues.
Figures


Similar articles
-
RNAi knockdown of dRNaseZ, the Drosophila homolog of ELAC2, impairs growth of mitotic and endoreplicating tissues.Insect Biochem Mol Biol. 2011 Mar;41(3):167-77. doi: 10.1016/j.ibmb.2010.12.001. Epub 2010 Dec 10. Insect Biochem Mol Biol. 2011. PMID: 21146608
-
Her2p molecular modeling, mutant analysis and intramitochondrial localization.Fungal Genet Biol. 2013 Nov;60:133-9. doi: 10.1016/j.fgb.2013.06.006. Epub 2013 Jul 11. Fungal Genet Biol. 2013. PMID: 23850602
-
Natural insertion of the bro-1 β-lactamase gene into the gatCAB operon affects Moraxella catarrhalis aspartyl-tRNA(Asn) amidotransferase activity.Microbiology (Reading). 2012 Sep;158(Pt 9):2363-2371. doi: 10.1099/mic.0.060095-0. Epub 2012 Jun 28. Microbiology (Reading). 2012. PMID: 22745266
-
Polo kinase and progression through M phase in Drosophila: a perspective from the spindle poles.Oncogene. 2005 Jan 10;24(2):230-7. doi: 10.1038/sj.onc.1208279. Oncogene. 2005. PMID: 15640838 Review.
-
Mitosis in Drosophila.J Cell Sci. 1989 Feb;92 ( Pt 2):137-46. doi: 10.1242/jcs.92.2.137. J Cell Sci. 1989. PMID: 2674166 Review.
Cited by
-
Biogenesis of glutaminyl-mt tRNAGln in human mitochondria.Proc Natl Acad Sci U S A. 2009 Sep 22;106(38):16209-14. doi: 10.1073/pnas.0907602106. Epub 2009 Sep 9. Proc Natl Acad Sci U S A. 2009. PMID: 19805282 Free PMC article.
-
Transcriptomic Changes of Bemisia tabaci Asia II 1 Induced by Chilli Leaf Curl Virus Trigger Infection and Circulation in Its Vector.Front Microbiol. 2022 Apr 28;13:890807. doi: 10.3389/fmicb.2022.890807. eCollection 2022. Front Microbiol. 2022. PMID: 35572639 Free PMC article.
-
The aminoacyl-tRNA synthetases of Drosophila melanogaster.Fly (Austin). 2015;9(2):53-61. doi: 10.1080/19336934.2015.1101196. Fly (Austin). 2015. PMID: 26761199 Free PMC article. Review.
-
Functional characterization of the Drosophila MRP (mitochondrial RNA processing) RNA gene.RNA. 2010 Nov;16(11):2120-30. doi: 10.1261/rna.2227710. Epub 2010 Sep 20. RNA. 2010. PMID: 20855541 Free PMC article.
-
Knockout of Drosophila RNase ZL impairs mitochondrial transcript processing, respiration and cell cycle progression.Nucleic Acids Res. 2015 Dec 2;43(21):10364-75. doi: 10.1093/nar/gkv1149. Epub 2015 Nov 8. Nucleic Acids Res. 2015. PMID: 26553808 Free PMC article.
References
-
- Altschul, S. F., W. Gish, W. Miller, E. W. Myers and D. J. Lipman, 1990. Basic local alignment search tool. J. Mol. Biol. 215 403–410. - PubMed
-
- Britton, J. S., W. K. Lockwood, L. Li, S. M. Cohen and B. A. Edgar, 2002. Drosophila's insulin/PI3-kinase pathway coordinates cellular metabolism with nutritional conditions. Dev. Cell 2 239–249. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

