Binding modes of CCR5-targetting HIV entry inhibitors: partial and full antagonists
- PMID: 18249144
- PMCID: PMC2701198
- DOI: 10.1016/j.jmgm.2007.12.003
Binding modes of CCR5-targetting HIV entry inhibitors: partial and full antagonists
Abstract
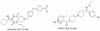
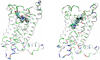
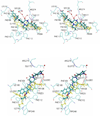
Since the CC-chemokine receptor 5 (CCR5) was identified as a major co-receptor for human immunodeficiency virus type 1 (HIV-1) entry into a host cell, CCR5-targetting HIV entry inhibitors have been developed and some of them are currently in clinical trials. Most of these inhibitors also inhibit the physiological chemokine reaction function of CCR5, which is so far considered to be safe to patients based on the observation that individuals that naturally lack CCR5 do not show apparent health problems. Nevertheless, to minimize the toxicity and side effects, it would be ideal to preserve the chemokine receptor activity. In this work, we simulated the flexible docking of two small molecule inhibitors to CCR5 in a solvated phospholipid bilayer environment. One of the inhibitors, aplaviroc has a unique feature of preserving two of the natural chemokine ligands binding to CCR5 and subsequent activation whereas the other one, SCH-C fully blocks chemokine-CCR5 interactions. Our results revealed significantly different binding modes of these two inhibitors although both established extensive interaction networks with CCR5. Comparison of the different binding modes suggests that avoiding the deep insertion of inhibitors into the transmembrane helix bundle may be able to preserve chemokine-CCR5 interactions. These results could help design HIV co-receptor activity-specific inhibitors.
Figures

Similar articles
-
HIV co-receptor CCR5: structure and interactions with inhibitors.Infect Disord Drug Targets. 2009 Jun;9(3):279-88. doi: 10.2174/1871526510909030279. Infect Disord Drug Targets. 2009. PMID: 19519482 Review.
-
Molecular interactions of CCR5 with major classes of small-molecule anti-HIV CCR5 antagonists.Mol Pharmacol. 2008 Mar;73(3):789-800. doi: 10.1124/mol.107.042101. Epub 2007 Dec 20. Mol Pharmacol. 2008. PMID: 18096812
-
Structure of CC Chemokine Receptor 5 with a Potent Chemokine Antagonist Reveals Mechanisms of Chemokine Recognition and Molecular Mimicry by HIV.Immunity. 2017 Jun 20;46(6):1005-1017.e5. doi: 10.1016/j.immuni.2017.05.002. Immunity. 2017. PMID: 28636951 Free PMC article.
-
Involvement of the second extracellular loop and transmembrane residues of CCR5 in inhibitor binding and HIV-1 fusion: insights into the mechanism of allosteric inhibition.J Mol Biol. 2008 Sep 12;381(4):956-74. doi: 10.1016/j.jmb.2008.06.041. Epub 2008 Jun 20. J Mol Biol. 2008. PMID: 18590744 Free PMC article.
-
Development of anti-HIV agents targeting dynamic supramolecular mechanism: entry and fusion inhibitors based on CXCR4/CCR5 antagonists and gp41-C34-remodeling peptides.Curr HIV Res. 2005 Oct;3(4):289-301. doi: 10.2174/157016205774370429. Curr HIV Res. 2005. PMID: 16250877 Review.
Cited by
-
Structural Insights to Human Immunodeficiency Virus (HIV-1) Targets and Their Inhibition.Adv Exp Med Biol. 2021;1322:63-95. doi: 10.1007/978-981-16-0267-2_3. Adv Exp Med Biol. 2021. PMID: 34258737
-
Structural insights from binding poses of CCR2 and CCR5 with clinically important antagonists: a combined in silico study.PLoS One. 2012;7(3):e32864. doi: 10.1371/journal.pone.0032864. Epub 2012 Mar 27. PLoS One. 2012. PMID: 22479344 Free PMC article.
-
Investigation of Inhibition Mechanism of Chemokine Receptor CCR5 by Micro-second Molecular Dynamics Simulations.Sci Rep. 2015 Aug 24;5:13180. doi: 10.1038/srep13180. Sci Rep. 2015. PMID: 26299310 Free PMC article.
-
Closing the door on flaviviruses: entry as a target for antiviral drug design.Antiviral Res. 2008 Oct;80(1):11-22. doi: 10.1016/j.antiviral.2008.05.004. Epub 2008 Jun 11. Antiviral Res. 2008. PMID: 18585795 Free PMC article. Review.
-
Pharmacotherapy of HIV-1 Infection: Focus on CCR5 Antagonist Maraviroc.Clin Med Ther. 2009;1:1497-1510. doi: 10.4137/cmt.s2365. Clin Med Ther. 2009. PMID: 19920876 Free PMC article.
References
-
- Deng H, Liu R, Ellmeier W, Choe S, Unutmaz D, Burkhart M, Di Marzio P, Marmon S, Sutton RE, Hill CM, Davis CB, Peiper SC, Schall TJ, Littman DR, Landau NR. Identification of a major co-receptor for primary isolates of HIV-1. Nature. 1996;381:661–666. - PubMed
-
- Wu L, Gerard NP, Wyatt R, Choe H, Parolin C, Ruffing N, Borsetti A, Cardoso AA, Desjardin E, Newman W, Gerard C, Sodroski J. CD4-induced interaction of primary HIV-1 gp120 glycoproteins with the chemokine receptor CCR-5. Nature. 1996;384:179–183. - PubMed
-
- Samson M, Libert F, Doranz BJ, Rucker J, Liesnard C, Farber CM, Saragosti S, Lapoumeroulie C, Cognaux J, Forceille C, Muyldermans G, Verhofstede C, Burtonboy G, Georges M, Imai T, Rana S, Yi Y, Smyth RJ, Collman RG, Doms RW, Vassart G, Parmentier M. Resistance to HIV-1 infection in caucasian individuals bearing mutant alleles of the CCR-5 chemokine receptor gene. Nature. 1996;382:722–725. - PubMed
-
- Liu R, Paxton WA, Choe S, Ceradini D, Martin SR, Horuk R, MacDonald ME, Stuhlmann H, Koup RA, Landau NR. Homozygous defect in HIV-1 coreceptor accounts for resistance of some multiply-exposed individuals to HIV-1 infection. Cell. 1996;86:367–377. - PubMed
-
- Cocchi F, DeVico AL, Garzino-Demo A, Arya SK, Gallo RC, Lusso P. Identification of RANTES, MIP-1 alpha, and MIP-1 beta as the major HIV-suppressive factors produced by CD8+ T cells. Science. 1995;270:1811–1815. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

