Histone H1 functions as a stimulatory factor in backup pathways of NHEJ
- PMID: 18250087
- PMCID: PMC2275134
- DOI: 10.1093/nar/gkn013
Histone H1 functions as a stimulatory factor in backup pathways of NHEJ
Abstract
DNA double-strand breaks (DSBs) induced in the genome of higher eukaryotes by ionizing radiation (IR) are predominantly removed by two pathways of non-homologous end-joining (NHEJ) termed D-NHEJ and B-NHEJ. While D-NHEJ depends on the activities of the DNA-dependent protein kinase (DNA-PK) and DNA ligase IV/XRCC4/XLF, B-NHEJ utilizes, at least partly, DNA ligase III/XRCC1 and PARP-1. Using in vitro end-joining assays and protein fractionation protocols similar to those previously applied for the characterization of DNA ligase III as an end-joining factor, we identify here histone H1 as an additional putative NHEJ factor. H1 strongly enhances DNA-end joining and shifts the product spectrum from circles to multimers. While H1 enhances the DNA-end-joining activities of both DNA Ligase IV and DNA Ligase III, the effect on ligase III is significantly stronger. Histone H1 also enhances the activity of PARP-1. Since histone H1 has been shown to counteract D-NHEJ, these observations and the known functions of the protein identify it as a putative alignment factor operating preferentially within B-NHEJ.
Figures




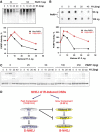
Similar articles
-
DNA ligase III as a candidate component of backup pathways of nonhomologous end joining.Cancer Res. 2005 May 15;65(10):4020-30. doi: 10.1158/0008-5472.CAN-04-3055. Cancer Res. 2005. PMID: 15899791
-
Involvement of poly(ADP-ribose) polymerase-1 and XRCC1/DNA ligase III in an alternative route for DNA double-strand breaks rejoining.J Biol Chem. 2004 Dec 31;279(53):55117-26. doi: 10.1074/jbc.M404524200. Epub 2004 Oct 21. J Biol Chem. 2004. PMID: 15498778
-
Requirement for Parp-1 and DNA ligases 1 or 3 but not of Xrcc1 in chromosomal translocation formation by backup end joining.Nucleic Acids Res. 2014 Jun;42(10):6380-92. doi: 10.1093/nar/gku298. Epub 2014 Apr 19. Nucleic Acids Res. 2014. PMID: 24748665 Free PMC article.
-
Backup pathways of NHEJ in cells of higher eukaryotes: cell cycle dependence.Radiother Oncol. 2009 Sep;92(3):310-5. doi: 10.1016/j.radonc.2009.06.024. Epub 2009 Jul 13. Radiother Oncol. 2009. PMID: 19604590 Review.
-
Mechanisms of DNA double strand break repair and chromosome aberration formation.Cytogenet Genome Res. 2004;104(1-4):14-20. doi: 10.1159/000077461. Cytogenet Genome Res. 2004. PMID: 15162010 Review.
Cited by
-
Gamma-H2AX in recognition and signaling of DNA double-strand breaks in the context of chromatin.Nucleic Acids Res. 2008 Oct;36(17):5678-94. doi: 10.1093/nar/gkn550. Epub 2008 Sep 4. Nucleic Acids Res. 2008. PMID: 18772227 Free PMC article. Review.
-
Role of H1 linker histones in mammalian development and stem cell differentiation.Biochim Biophys Acta. 2016 Mar;1859(3):496-509. doi: 10.1016/j.bbagrm.2015.12.002. Epub 2015 Dec 13. Biochim Biophys Acta. 2016. PMID: 26689747 Free PMC article. Review.
-
An in vitro DNA double-strand break repair assay based on end-joining of defined duplex oligonucleotides.Methods Mol Biol. 2012;920:485-500. doi: 10.1007/978-1-61779-998-3_33. Methods Mol Biol. 2012. PMID: 22941624 Free PMC article.
-
DNA double-strand break repair as determinant of cellular radiosensitivity to killing and target in radiation therapy.Front Oncol. 2013 May 10;3:113. doi: 10.3389/fonc.2013.00113. eCollection 2013. Front Oncol. 2013. PMID: 23675572 Free PMC article.
-
Optimal function of the DNA repair enzyme TDP1 requires its phosphorylation by ATM and/or DNA-PK.EMBO J. 2009 Dec 2;28(23):3667-80. doi: 10.1038/emboj.2009.302. Epub 2009 Oct 22. EMBO J. 2009. PMID: 19851285 Free PMC article.
References
-
- Khanna KK, Jackson SP. DNA double-strand breaks: signaling, repair and the cancer connection. Nat. Genet. 2001;27:247–254. - PubMed
-
- Sancar A, Lindsey-Boltz LA, Ünsal-Kacmaz K, Linn S. Molecular mechanisms of mammalian DNA repair and the DNA damage checkpoints. Annu. Rev. Biochem. 2004;73:39–85. - PubMed
-
- Ahnesorg P, Smith P, Jackson SP. XLF interacts with the XRCC4-DNA Ligase IV complex to promote DNA nonhomologous end-Joining. Cell. 2006;124:301–313. - PubMed
-
- Buck D, Malivert L, de Chasseval R, Barraud A, Fondaneche M-C, Sanal O, Plebani A, Stephan J-L, Hufnagel M, le Deist F. Cernunnos, a novel nonhomologous end-joining factor, is mutated in human immunodeficiency with microcephaly. Cell. 2006;124:287–299. - PubMed
-
- Nevaldine B, Longo JA, Hahn PJ. The scid defect results in much slower repair of DNA double-strand breaks but not high levels of residual breaks. Radiat. Res. 1997;147:535–540. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

