Structure/function analysis of yeast ribosomal protein L2
- PMID: 18263608
- PMCID: PMC2330241
- DOI: 10.1093/nar/gkn034
Structure/function analysis of yeast ribosomal protein L2
Abstract
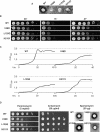
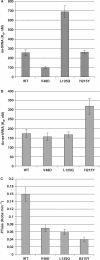
Ribosomal protein L2 is a core element of the large subunit that is highly conserved among all three kingdoms. L2 contacts almost every domain of the large subunit rRNA and participates in an intersubunit bridge with the small subunit rRNA. It contains a solvent-accessible globular domain that interfaces with the solvent accessible side of the large subunit that is linked through a bridge to an extension domain that approaches the peptidyltransferase center. Here, screening of randomly generated library of yeast RPL2A alleles identified three translationally defective mutants, which could be grouped into two classes. The V48D and L125Q mutants map to the globular domain. They strongly affect ribosomal A-site associated functions, peptidyltransferase activity and subunit joining. H215Y, located at the tip of the extended domain interacts with Helix 93. This mutant specifically affects peptidyl-tRNA binding and peptidyltransferase activity. Both classes affect rRNA structure far away from the protein in the A-site of the peptidyltransferase center. These findings suggest that defective interactions with Helix 55 and with the Helix 65-66 structure may indicate a certain degree of flexibility in L2 in the neck region between the two other domains, and that this might help to coordinate tRNA-ribosome interactions.
Figures

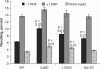
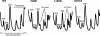

Similar articles
-
Yeast ribosomal protein L10 helps coordinate tRNA movement through the large subunit.Nucleic Acids Res. 2008 Nov;36(19):6187-98. doi: 10.1093/nar/gkn643. Epub 2008 Sep 29. Nucleic Acids Res. 2008. PMID: 18824477 Free PMC article.
-
The central core region of yeast ribosomal protein L11 is important for subunit joining and translational fidelity.Mol Genet Genomics. 2011 Jun;285(6):505-16. doi: 10.1007/s00438-011-0623-2. Epub 2011 Apr 26. Mol Genet Genomics. 2011. PMID: 21519857 Free PMC article.
-
Optimization of ribosome structure and function by rRNA base modification.PLoS One. 2007 Jan 24;2(1):e174. doi: 10.1371/journal.pone.0000174. PLoS One. 2007. PMID: 17245450 Free PMC article.
-
The social life of ribosomal proteins.FEBS J. 2005 May;272(9):2098-108. doi: 10.1111/j.1742-4658.2005.04651.x. FEBS J. 2005. PMID: 15853795 Review.
-
Pseudouridines and pseudouridine synthases of the ribosome.Cold Spring Harb Symp Quant Biol. 2001;66:147-59. doi: 10.1101/sqb.2001.66.147. Cold Spring Harb Symp Quant Biol. 2001. PMID: 12762017 Review.
Cited by
-
Nervous-Like Circuits in the Ribosome Facts, Hypotheses and Perspectives.Int J Mol Sci. 2019 Jun 14;20(12):2911. doi: 10.3390/ijms20122911. Int J Mol Sci. 2019. PMID: 31207893 Free PMC article. Review.
-
The role of ribosomal protein networks in ribosome dynamics.Nucleic Acids Res. 2025 Jan 7;53(1):gkae1308. doi: 10.1093/nar/gkae1308. Nucleic Acids Res. 2025. PMID: 39788545 Free PMC article.
-
A molecular clamp ensures allosteric coordination of peptidyltransfer and ligand binding to the ribosomal A-site.Nucleic Acids Res. 2010 Nov;38(21):7800-13. doi: 10.1093/nar/gkq641. Epub 2010 Jul 21. Nucleic Acids Res. 2010. PMID: 20660012 Free PMC article.
-
Ribosomes in the balance: structural equilibrium ensures translational fidelity and proper gene expression.Nucleic Acids Res. 2014 Dec 1;42(21):13384-92. doi: 10.1093/nar/gku1020. Epub 2014 Nov 11. Nucleic Acids Res. 2014. PMID: 25389262 Free PMC article.
-
Tracking fluctuation hotspots on the yeast ribosome through the elongation cycle.Nucleic Acids Res. 2017 May 5;45(8):4958-4971. doi: 10.1093/nar/gkx112. Nucleic Acids Res. 2017. PMID: 28334755 Free PMC article.
References
-
- Inoue T, Sullivan FX, Cech TR. Intermolecular exon ligation of the rRNA precursor of Tetrahymena: oligonucleotides can function as 5′ exons. Cell. 1985;43:431–437. - PubMed
-
- Kazemie M. Binding of aminoacyl-tRNA to reconstituted subparticles of Escherichia coli large ribosomal subunits. Eur. J. Biochem. 1976;67:373–378. - PubMed
-
- Fahnestock SR. Evidence of the involvement of a 50S ribosomal protein in several active sites. Biochemistry. 1975;14:5321–5327. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

