Chromatin structure and expression of a gene transcribed by RNA polymerase III are independent of H2A.Z deposition
- PMID: 18268003
- PMCID: PMC2293117
- DOI: 10.1128/MCB.01953-07
Chromatin structure and expression of a gene transcribed by RNA polymerase III are independent of H2A.Z deposition
Abstract
The genes transcribed by RNA polymerase III (Pol III) generally have intragenic promoter elements. One of them, the yeast U6 snRNA (SNR6) gene is activated in vitro by a positioned nucleosome between its intragenic box A and extragenic, downstream box B separated by approximately 200 bp. We demonstrate here that the in vivo chromatin structure of the gene region is characterized by the presence of an array of positioned nucleosomes, with only one of them in the 5' end of the gene having a regulatory role. A positioned nucleosome present between boxes A and B in vivo does not move when the gene is repressed due to nutritional deprivation. In contrast, the upstream nucleosome which covers the TATA box under repressed conditions is shifted approximately 50 bp further upstream by the ATP-dependent chromatin remodeler RSC upon activation. It is marked with the histone variant H2A.Z and H4K16 acetylation in active state. In the absence of H2A.Z, the chromatin structure of the gene does not change, suggesting that H2A.Z is not required for establishing the active chromatin structure. These results show that the chromatin structure directly participates in regulation of a Pol III-transcribed gene under different states of its activity in vivo.
Figures
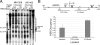
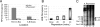
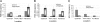
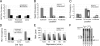
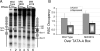

References
-
- Albert, I., T. N. Mavrich, L. P. Tomsho, J. Qi, S. J. Zanton, S. C. Schuster, and B. F. Pugh. 2007. Translational and rotational settings of H2A.Z nucleosomes across the Saccharomyces cerevisiae genome. Nature 446572-576. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
