De novo high-resolution protein structure determination from sparse spin-labeling EPR data
- PMID: 18275810
- PMCID: PMC2390841
- DOI: 10.1016/j.str.2007.11.015
De novo high-resolution protein structure determination from sparse spin-labeling EPR data
Abstract
As many key proteins evade crystallization and remain too large for nuclear magnetic resonance spectroscopy, electron paramagnetic resonance (EPR) spectroscopy combined with site-directed spin labeling offers an alternative approach for obtaining structural information. Such information must be translated into geometric restraints to be used in computer simulations. Here, distances between spin labels are converted into distance ranges between beta carbons by using a "motion-on-a-cone" model, and a linear-correlation model links spin-label accessibility to the number of neighboring residues. This approach was tested on T4-lysozyme and alphaA-crystallin with the de novo structure prediction algorithm Rosetta. The results demonstrate the feasibility of obtaining highly accurate, atomic-detail models from EPR data by yielding 1.0 A and 2.6 A full-atom models, respectively. Distance restraints between amino acids far apart in sequence but close in space are most valuable for structure determination. The approach can be extended to other experimental techniques such as fluorescence spectroscopy, substituted cysteine accessibility method, or mutational studies.
Figures





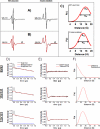
References
-
- Mathematica Champaign. Wolfram Research, Inc; Illinois: 2005.
-
- Altenbach C, Oh KJ, et al. Estimation of inter-residue distances in spin labeled proteins at physiological temperatures: experimental strategies and practical limitations. Biochemistry. 2001;40(51):15471–82. - PubMed
-
- Baker D. A surprising simplicity to protein folding. Nature. 2000;405(6782):39–42. - PubMed
-
- Berman HM, Battistuz T, et al. The Protein Data Bank. Acta Crystallogr D Biol Crystallogr. 2002;58(Pt 6 No 1):899–907. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

